What makes the body absorb too much iron?
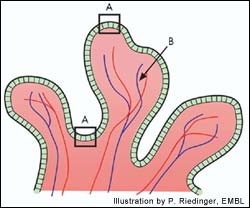
Cells in the intestine (A) absorb iron that is passed along to the bloodstream (B). Previously it was thought that the Hfe mutation found in Hemochromatosis acted to change the behaviour of these cells. The latest research shows that the mutation seemsto act primarily in the liver.
Researchers at EMBL and Harvard gain new insights into hemochromatosis
Like most nutrients, iron is good for people – in the right doses. When the body has enough iron, our cells stop absorbing it from food; if there is too little, they absorb more. This system breaks down in the most common inherited disease in the Western world: hemochromatosis, which affects about one in every 250 people and is often fatal if it is not recognized and treated. Now researchers at the European Molecular Biology Laboratory in Heidelberg (EMBL) and Harvard Medical School (U.S.) have linked the response of a gene in the liver to the disease. The study, which appears in the current issue of Nature Genetics, is changing our understanding of how hemochromatosis develops.
“Untreated iron overload can result in liver cancer, heart disease, or other fatal conditions,” says Martina Muckenthaler, a staff scientist at EMBL in the research group of Matthias Hentze. “The mutation that causes the disease was thought to have its major effects in the intestine, where cells absorb iron from food. Our current study has changed that picture.”
Sufferers of hemochromatosis have inherited a defective copy of a gene called Hfe from each parent. The mutation can be traced back several centuries into Celtic history, where it originated in a single person who passed it down to his or her children. It has now spread throughout Europe and the Western world.
The defect leads a person’s intestine to absorb too much iron from his or her diet. Over the course of many years, this builds up, and by middle age, the overload may be very serious. Hentze and his group at EMBL have been trying to trace these effects back to their molecular causes.
Until now, most researchers have thought that the problem stems from a faulty intestinal signal that there is too little iron, leading cells in the intestines to produce more transport proteins that draw iron inside.
By combining two approaches, the EMBL and Harvard teams have now rewritten this model. Their study shows that the link between Hfe and iron is probably in the liver, rather than the intestine.
The EMBL group was investigating iron absorption with a technology called DNA chips. The method allows scientists to monitor how a cell’s genes respond to changes – such as a raise in levels of iron. Working with the research team of Wilhelm Ansorge, Muckenthaler put together a set of probes that could watch for changes in 300 genes known to have some connection to iron.
Simultaneously, Nancy Andrews’ group at Harvard was working on hemochromatosis in mice – which are often used to create models of human disease. The Harvard scientists had developed strains of mice without Hfe, or with the mutant version found in hemochromatosis. Both strains absorbed too much iron and exhibited the symptoms of the disease.
The DNA chips were used to compare cells taken from these mice and their healthy cousins. In the former model, cells with defective Hfe were expected to behave like those of healthy mice with an iron deficiency – they ought to produce more iron-absorbing proteins in the intestine. That wasn’t the case.
So the researchers started to look farther afield for other effects of mutant Hfe. They discovered something intriguing in the liver. A molecule called hepcidin wasn’t being activated properly.
When iron levels rise in a healthy mouse, its liver produces more hepcidin, and hepcidin acts like a hormone to reduce iron absorption by the intestine. This didn’t happen in mice with mutant or missing Hfe – even when the researchers directly injected them with iron. This was proof that Hfe is necessary to make a connection between iron and hepcidin, and that shakes up the old model. The importance of low hepcidin production in hemochromatosis is further underlined by the work of Sophie Vaulont’s team at the Cochin Institute in Paris, which appears in the same issue of Nature Genetics. They were able to show that the liver does not load iron when hepcidin is added to mice lacking the hemochromatosis gene. It means that focus should change from the intestine to the liver – where the lack of Hfe seems to be doing the most damage.
How can hepcidin expression by the liver change iron absorption among intestinal cells? A lot of questions remain, Muckenthaler says, but data from the DNA chips suggest how this might be happening. Without hepcidin, the body can’t seem to put the brakes on intestinal molecule, called Cybrd1.
Normally iron atoms in that form they can’t be brought into cells. Cybrd1’s job is to reduce the number of charges to two, making iron transportable. If it were active all the time, a lot more of the body’s iron would be absorbable. When it isn’t active, there is very little to be absorbed. Hepcidin’s normal function might be to lock down Cybrd1, thereby shutting down the supply of available iron. This would make activating hepcidin – which can’t happen with a mutant form of Hfe – essential to stopping the flow of iron into the body.
This spells an important shift in hemochromatosis research: scientists will now turn their attention to the liver, in hopes of working out the details of the circuit that connects iron, Hfe and, hepcidin. It also hints strongly at a link between the body’s uptake of iron and the immune system, where hepcidin also has a role to play.
Source article
Regulatory defects in liver and intestine implicate abnormal hepcidin and Dcytb expression in mouse hemochromatosis.
Martina Muckenthaler, Cindy N.Roy, Ángel O. Custodio, Belén Miñana, Jos deGraaf, Lynne K.Montross,Nancy C.Andrews and Matthias W. Hentze
Nature Genetics, 21 April 2003, dx.doi.org/doi:10.1038/Ng1152
Scientific contacts
Matthias Hentze
EMBL
Meyerhofstrasse 1
D–69117 Heidelberg
Germany
Tel: +49 6221 387 501 or 502
Fax: +49 6221 387518
E-mail: hentze@embl.de
Martina Muckenthaler
EMBL
Meyerhofstrasse 1
D–69117 Heidelberg
Germany
Tel: +49 6221 387 502 or 410
Fax: +49 6221 387518
E-mail: muckenthaler@embl.de
Media Contact
More Information:
http://www.embl-heidelberg.de/ExternalInfo/oipa/All latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles
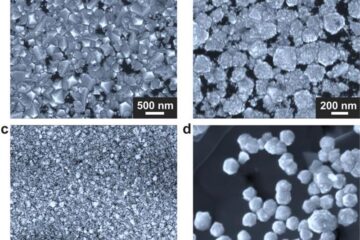

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…