Researchers map minority microbes in the colon

In a new study, researchers take a first up close look at these “hydrogenotrophic” microbes, mapping where they live and how abundant they are in different parts of the lower intestine.
The findings are reported in the International Society for Microbial Ecology Journal.
This is the first study to sample these – or any other – microbes at specific locales in the colon, said University of Illinois animal sciences and Institute for Genomic Biology professor Rex Gaskins, who led the research with Carle Foundation gastroenterologist Dr. Eugene Greenberg. These organisms are particularly difficult to get at because they inhabit a thick layer of protective mucous that lines the colon. Previous studies looked only at microbes passed in stool, Gaskins said.
“With those methods, the less abundant taxa were often undetected,” he said. “Little was known about their ecology, or the extent to which the mucosa is colonized and what kind of variation exists.”
This is also the first study to compare the diversity and abundance of colonic microbes in multiple (in this case 25) healthy individuals, he said.
Scientists have known for decades that various microbes in the colon collaborate to ferment undigested food and degrade and dispose of the byproducts of fermentation. (M.P. Bryant, who was an animals sciences professor at Illinois, discovered the first hydrogen-based symbiotic relationship among microbes of the gut in the 1960s.)
Disruptions of the colonic ecosystem can have profound implications for human health, Greenberg said. Although there is a genetic component as well, evidence suggests that inflammatory bowel disease (which includes Crohn’s disease and ulcerative colitis) emerges in response to a microbial imbalance, he said. A study led by Gaskins in 2006 demonstrated for the first time that hydrogen sulfide, a major byproduct of bacterial fermentation in the gut, is genotoxic (damaging to DNA) and may lead to colon cancer.
“We’re getting closer and closer to looking at the microbiological origin of many diseases,” Greenberg said.
To get a better picture of the microbes in colonic mucosa, the researchers analyzed biopsies obtained from healthy subjects during routine colonoscopy exams. They sampled the right (ascending) and left (descending) colon as well as the rectum, then looked for genes that contribute to different hydrogen-consuming metabolic pathways.
The researchers found that all of the subjects harbored three important classes of hydrogen-consuming microbes: methanogens, which convert hydrogen to methane; acetogens, which make acetate from carbon dioxide and hydrogen; and sulfate-reducing bacteria, which expel hydrogen sulfide gas.
Methanogens accounted for about half of the microbes seen in each region of the colon and “become more abundant as you advance toward the rectum,” said postdoctoral researcher Franck Carbonero, an author on the study.
Sulfate-reducing bacteria (SRB) thrive in the presence of sulfate (a byproduct of eating foods such as meat, milk or eggs that have high levels of sulfur-containing amino acids).
The SRB generally outnumbered acetogens in the right colon, Carbonero said, while acetogens were more abundant than SRB in the left colon and rectum.
The researchers found variation among individual study subjects, particularly in the abundance of the three classes of microbes at different biopsy sites. And the team made a surprising discovery when analyzing biopsies taken less than a centimeter apart: microbial diversity can occur even at this small scale.
“These data indicate that if you get down to the scale at which these microbes make their living, perhaps it’s not all the same,” Gaskins said. “We haven’t begun yet to think at that spatial scale.”
The finding is relevant because disease tends to originate in very specific regions of the colon, Greenberg said. Ulcerative colitis starts at the rectum and then proceeds “upriver,” he said. “When you see Crohn’s disease you see an area of involvement, then it’s normal and then you see another area of involvement. And if you do surgery and remove the disease, the disease almost always recurs from the point of removal because – we believe – it’s been reseeded with microbes.”
Future studies will examine the role of hydrogenotrophic microbes in constipation and will look at how diet affects the composition and abundance of microbes in the colon, the researchers said.
The Carle Foundation and the Carle Hospital-University of Illinois Translational Research Program supported this research.
Editor’s notes: To reach Rex Gaskins, call 217- 244-3165; email hgaskins@illinois.edu.
The paper, “Abundance and Diversity of Mucosa-Associated Hydrogenotrophic Microbes in the Healthy Human Colon,” is available online or from the U. of I. News Bureau
Media Contact
More Information:
http://www.illinois.eduAll latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles
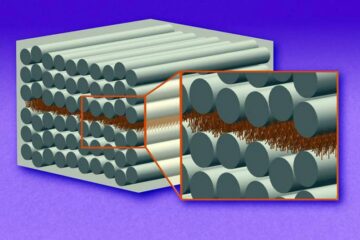

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
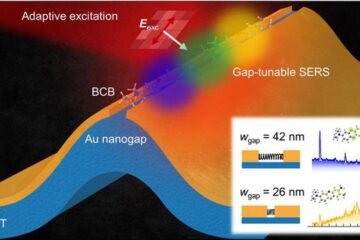
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…