3-D model reveals secrets of metastasis

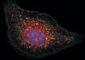
Working in the labs of Whitehead Member Paul Matsudaira and MIT professor Douglas Lauffenburger, postdoctoral researcher Muhammad Zaman discovered that cells move quite differently in three dimensions. His study, which focused on human prostate tumor cells, appeared this week in the online early edition of Proceedings of the National Academy of Sciences.
“Two-dimensional assays ignore the obstacles that cells face in their natural contexts,” explains Zaman, who recently became an assistant professor at the University of Texas at Austin. “In 3D, cells move through a thick jungle of fibers, or “vines”TM, that hinder forward progress.”
Cells must either squeeze through or chop up these putative vines to get anywhere. As a result, they move slower in three dimensions.
In an interesting twist, all cells need at least some vines to move, as they latch onto the “branches” with claw-like proteins called integrins and pull themselves forward. When Zaman disabled some of these claws, in a manner analogous to certain anti-cancer drugs, the cells moving across the top of the jungle canopy (in two dimensions) needed a greater number of vines to keep up their pace, while cells plowing through the jungle instead needed vines chopped to maintain the same speed. The complexity of this situation is further increased in that the cells become dramatically sensitive to the stiffness of the vines when the integrins are disabled and consequently tend to squeeze through the vines rather than pushing them aside.
“Our findings help explain why two-dimensional assays for metastasis-inhibiting drugs do not effectively predict their effects in tissue,” says Lauffenburger, who is director of MITâ€TMs Biological Engineering Division. He believes pharmaceutical companies will eventually use three-dimensional assays, accompanied by appropriate computational models such as that also recently published by Zaman (in Biophysical Journal in 2005), to determine how drugs affect metastasis.
But technology must improve before more complicated 3D studies are attempted. For his 3D work Zaman worked with one sample at a time, using a special confocal microscope at the Whitehead-MIT BioImaging Center. The microscope divided each specimen into virtual slices, generating a new stack of images every 15 minutes.
“It took me about a year to get enough data because the microscope wasnâ€TMt designed for high-throughput experiments,” he says. Fortunately, the BioImaging Center has one of the most powerful sets of computers at MIT and the imaging processing and analysis went quite quickly.
“Muhammad was successful for two reasons,” says Matsudaira. “His computational model predicted what would happen in virtual experiments and then he was able to go straight to test the predictions with these complicated 3D experiments. As a result, the sophisticated models of cell movement enhance our understanding of key biological processes, including metastasis.”
Media Contact
More Information:
http://www.wi.mit.eduAll latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…