Biochips in Drug Development
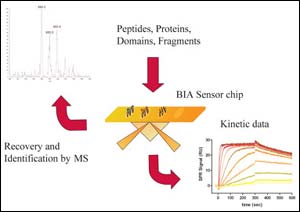
Figure 1: Ligand fishing (BIA-MS coupling)
Biomolecular interaction analysis (BIA) using SPR (surface plasmon resonance) biosensors is now utilised increasingly in nearly all phases of drug development. The BIA system consists out of a light source emitting near infrared light, a sensor microchip, an automated liquid handling system with constant flow and a diode array position-sensitive detector. One of the two interacting partners (referred to as the ligand) is immobilized on the sensor surface. The other binding partner, called the analyte, is directed over the surface in a constant flow system allowing to monitor the interaction of the binding partners in “real time”.
BIA covers a broad range of applications in target identification and validation, secondary screens and lead optimisation, early ADME assays as well as testing immunoreactivity.
Assay development and functional assays of proteins are the most commonly used areas of applications of BIA in drug development.
Advantages of SPR-based measurements are the possibility to measure in real time without the need of labelling yielding highly reproducible kinetic data with low sample consumption.
Ligand fishing
BIA-technology is ideally suited as a micro-affinity purification platform allowing on-line detection of binding events and the direct quantification of bound material on biological surfaces of interest.
Kinetic parameters may be obtained as additional information. Provided that a binding partner has been captured on the sensor chip it can be recovered by a gentle elution for further downstream analyses like mass spectrometry or western blot analysis (see figure 1).
In comparison to classical chromatographic methods biosensor systems show reduced unspecific binding, are compatible with small volume recovery and are readily automated.
The identification of unknown ligands is of particular interest for revealing function of orphan receptors, for the detection of cellular interaction networks and for testing the biocompatibility of novel surface coatings.
Target validation
Genomics and proteomics based techniques have provided a lot of new potential targets which has to be validated to prove that a DNA, RNA or protein molecule is directly involved in a disease process and is therefore a suitable target for the development of new therapeutic compounds.
The assessment of biological function, involvement in biological pathways and role in pathogenesis of potential targets can be achieved by mapping and validation of protein interaction networks in vivo and in vitro.
Yeast two hybrid and phage display are classical approaches for systematic protein interaction screening capable to probe millions of interactions. Positives obtained from these screening technologies can be validated using BIA technology.
Furthermore, additional parameters relevant for the interaction of interest can be investigated. This could be interactions with multiple components, effects of cofactors, pH-changes or the role of posttranslational modifications.
Lead optimisation
Hit validation is needed to determine whether a molecule identified in a screen or assay will eventually lead to a drug.
Therefore secondary assays generating data about potency, selectivity and functional biochemical activity have to be performed.
BIA technology has an enormous capability for the rapid confirmation of hits from high throughput screens by a comprehensive kinetic characterisation of potential lead compounds.
Information about affinities, rates of association and dissociation in complex formation and binding stoichiometries is very valuable for a ranking and optimisation of lead compounds. Even compounds binding with low affinity or transient kinetics, often found in early phases of drug development, can accurately be identified.
The kinetic information obtained by functional assays together with the knowledge about structural properties of biomolecules allows predictions on structure activity relationships.
Lead optimisation is an iterative process involving computer-assisted molecular modelling, chemical synthesis of new compounds and functional assays.
BIA technology is used in lead optimisation by linking compound structural information with a comprehensive kinetic characterisation of ligand binding.
ADME
ADME assays (adsorption, distribution, metabolism, elimination) become more and more important even in the earlier phases of the drug development process.
BIA assays provide very valuable information for a cost-effective in vitro characterisation of potential drug candidates in early ADME studies, although there are no methods available for accurately predicting what will happen to a drug in vivo.
SPR biosensor technology has already been applied for the binding of potential drug candidates to serum proteins, for analysing the adsorption of small molecules to artificial membranes immobilised on sensor chips and for measuring the influence of compounds on metabolic pathways.
The pharmacokinetic properties of small molecule drugs are a function of the reversible binding to serum proteins such as serum albumin, alpha 1-acid glycoprotein or gamma globulins, reducing the bioavailibility of the drug (see figure 2).
Biaffin offers a BIA assay for analysing the binding of small molecule compounds to high density serum protein surfaces yielding valuable information about affinitiy, solubility and binding stoichiometry of potential drug candidates.
Immune response
Newly designed pharmaceuticals can cause an unwanted immune response. In preclinical studies such potentials can be recognised in animal studies.
BIA technology can detect antibodies against new drugs in animal serum samples. With appropriate control experiments positive binding signals of low antibody levels can clearly be identified as a specific signal despite of the complexity of the protein mixture in serum samples.
Media Contact
All latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…