Flu tracked to viral reservoir in tropics

“We now know where the influenza A virus comes from every year,” said Edward Holmes, professor of biology at Penn State. “And because we now know how the virus evolves, we have a much better chance of controlling it.”
Currently, there are many strains of the influenza virus that appear only in birds, which are natural viral reservoirs. So far three of these viral strains — H1N1, H2N2 and H3N2 – have caused epidemics in humans as influenza A.
Of the three, H3N2 is the dominant strain, responsible for most influenza infections each winter, with lower levels of H1N1. However, little is known about how these two strains spread on a geographical scale, and how whole genome of influenza A virus evolves.
Holmes and his colleagues analyzed complete genomes of 1,032 strains of H1N1 and H3N2 viruses sampled over a 12-year period from New York state in the northern hemisphere and New Zealand in the southern hemisphere.
The researchers noticed that over time, both strains follow a distinctive pattern. In seasons where the H3N2 strain is dominant, H1N1 is not and vice versa.
“We found that the two strains peak at different times, and seem to be directly competing with each other” said Holmes, whose findings appear today online in Nature. The results also indicate that compared to the H3N2 strain, the H1N1 strain exhibits far less genetic diversity, although it is not clear why.
Holmes says his results also show that the influenza A virus is frequently exchanging genes by reassortment – when multiple human influenza viruses infect a single person and shuffle their genes – which sometimes allows the virus to acquire a new haemagglutinin, a protein that facilitates the entry of viral particles into the host cells.
These new haemagglutinins sometimes cause vaccines to fail, explained Holmes, whose work is funded by the National Institutes of Health.
“The critical thing is unless you understand the way the genome evolves, you will not understand why vaccines work during some years and fail during others,” he added. “We can now show that vaccines failed in some years because new haemagglutinins appeared.”
The Penn State researcher says his analysis not only indicates how the influenza virus is evolving, but also where new strains are being generated.
Each year new strains appear in the northern hemisphere, infect people and then burn out. However, patterns of genetic diversity within the viruses suggest the strains are coming from a global source population. The researchers believe that there must be some reservoir somewhere that every year generates new strains that are injected each season into the north and the south, and then burn themselves out.
“We know the strains are dying out every year in the northern and southern hemispheres. So they're surviving somewhere else, and we think it is a reservoir in the tropics,” Holmes said. “It tells us that to really understand how the influenza virus evolves on a seasonal basis, and to make the best vaccine, we need to focus our surveillance on the source population in the tropics, especially in places such as Southeast Asia.”
Media Contact
More Information:
http://www.psu.eduAll latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles
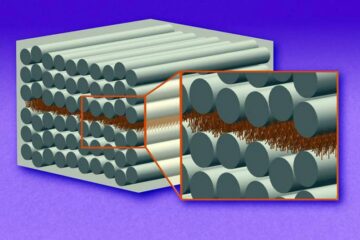

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
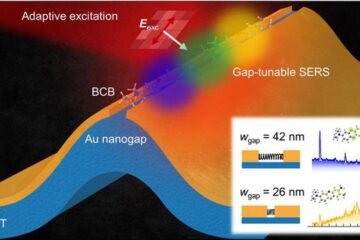
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…