Advance Cuts Brain Mapping Time from Years to Months

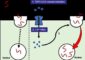
Mapping the billions of connections in the brain is a grand challenge in neuroscience. The current method for mapping interconnected brain cells involves the use of room-size microscopes known as transmission electron microscopes (TEMs).
Until now the process of mapping even small areas of the brain using these massive machines would have required several decades. In this week's open-access journal PLoS Biology, research teams at the University of Utah John A. Moran Eye Center and the University of Colorado at Boulder report technical advances that have reduced the time it takes to process high-speed “color” ultrastructure mapping of brain regions down to a few months.
These advances did not require the invention of new electron microscopes. The technical leap comes mostly from new powerful software that “takes over” the building, connecting and viewing of terabyte scale pictures produced by TEMs. Perhaps just as important is the fact that these researchers are now making these technologies available worldwide to scientists in multiple fields of research. “Our goals were to unleash a global network of electron microscopes and provide web-accessible imagery for battalions of brain network analysts,” said Robert Marc, Ph.D., Director of Research for the Moran Eye Center at the University of Utah.
Marc and this team of researchers have been working on this project for eight years. They have refined the software and molecular tools to where they are now able to share them globally. “This changes the playing field for building brain maps from a few specialized laboratories to the desktops of biologists world-wide,” says Marc.
The automation tools developed at the University of Colorado at Boulder, Center for 3D Electron Microscopy, allow capture of 25,000 TEM images weekly. At the same time, the Scientific Computing and Imaging Institute at the University of Utah developed software to automatically merge thousands of images into gigabyte-scale mosaics and align the mosaics into terabyte-scale volumes. And in parallel, teams at the Moran Eye Center developed TEM-compatible molecular probes and classification software to tag every cell with a molecular signature, creating “color” TEM imaging.
As part of this project, the authors will soon reveal the very first molecular level map of the entire retina and neuronal networks in both a normal mammalian retina and genetic models of retinal degeneration. More than 92 percent of this 400,000-image volume has been built and should be complete by mid-April. Lead author James Anderson explains, “This technology lets the neuroscience field build circuitry blueprints for healthy neural tissues. We can compare diseased tissues to these blueprints to understand how they rewire the brain and use them to evaluate the effectiveness of treatments which aren't detectable with other methods.”
While this molecular model of the retina is groundbreaking, the technology will extend far beyond ophthalmology. This is a new way to explore traumatic brain injury, neurodegenerative diseases, and epilepsy. It also makes screening of genetic models practical. By combining these tools with advanced image processing, the questions that can be explored by TEM may be limited only by the imaginations of biologists.
Important Notes for Members of the Media
PLEASE ADD THE LINK TO THE PUBLISHED ARTICLE IN ONLINE VERSIONS OF YOUR REPORT: http://biology.plosjournals.org/perlserv/?request=get-document&doi=10.1371/journal.pbio.1000074
All works published in PLoS Biology are open access. Everything is immediately available—to read, download, redistribute, include in databases, and otherwise use—without cost to anyone, anywhere, subject only to the condition that the original authorship and source are properly attributed. Copyright is retained by the authors. The Public Library of Science uses the Creative Commons Attribution License.
Media Contact
More Information:
http://www.utah.eduAll latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles

Wildfire danger to increase due to climate change
WSL Institute for Snow and Avalanche Research (SLF) researchers expect an elevated wildfire danger in the Alpine Foreland from 2040 onwards due to changing meteorological conditions. The danger currently remains…
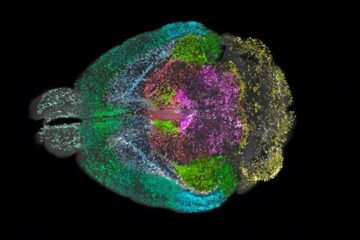
Advanced Brain Science Without Coding Expertise
Researchers at Helmholtz Munich and the LMU University Hospital Munich introduce DELiVR, offering a new AI-based approach to the complex task of brain cell mapping. The deep learning tool democratizes…
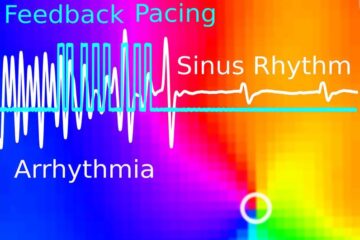
Gentle defibrillation for the heart
Using light pulses as a model for electrical defibrillation, Göttingen scientists developed a method to assess and modulate the heart function. The research team from the Max Planck Institute for…