New imaging technique tracks cancer-killing cells over prolonged period

Now, for the first time, researchers at the Stanford University School of Medicine have devised a way to obtain repeated “snapshots” of the location and survival of such cells in a living human patient months and possibly years later. This is good news for individual patients and clinicians who may want to assess the cells' disease-fighting performance over time, as well as for researchers trying to design more effective cell-based therapies.
“This has never before been done in a human,” said the senior author of the research, Sanjiv Gambhir, MD, PhD, director of Stanford's Molecular Imaging Program. “Until now, we've been shooting blind—never knowing why failed therapies didn't work. Did the cells die? Did they not get where we wanted them to go? Now we can repeatedly monitor them throughout their lifetime.” Gambhir is a professor of radiology and a member of Stanford's Cancer Center. He collaborated with researchers at City of Hope in Los Angeles and at UCLA to conduct the research.
Gambhir and his colleagues tested the technique in a middle-aged man with an aggressive brain tumor (called a glioblastoma) who was enrolled in a clinical trial of cell-based therapy at City of Hope. However, they believe similar strategies will work to monitor cell-based therapies for many disorders. The results of the case study will be published online Nov. 18 in Nature Clinical Practice Oncology.
The new approach relies on a two-step process: first, the therapeutic cells are modified to express a unique reporter gene shared by no other cells in the body. Second, an imaging agent that is trapped only in cells expressing the reporter gene is injected into the patient. The unbound imaging agent is otherwise quickly cleared from the body. Each time the imaging agent is used, the researchers get a new, up-to-date map showing the cells' locations.
The technique has several advantages over previous tracking methods. Unlike an external radioactive tag, which decays over a short time and does not indicate whether a cell is living or dead, the reporter gene is expressed throughout a cell's lifetime, but not beyond. Furthermore, unlike an external tag, the reporter gene is duplicated and passed along if the original cell divides. Finally, different reporter genes can be used that could indicate not only the location of the cells, but also what they're up to.
“In this patient, the reporter gene was always on,” said Gambhir. “But the beauty of this approach is that we could make it so the reporter gene is expressed only if the cell differentiates, or finds a certain target. Has the T cell found a tumor? Has it activated its cell-killing machinery?”
In the current study, Gambhir collaborated with Michael Jensen, MD, associate chair of the cancer immunotherapeutics & tumor immunology program at City of Hope, and others to remove cytotoxic, or “killer,” T cells from the man with glioblastoma. These cells naturally seek out and destroy infected or dysfunctional cells in the body. The researchers then inserted a circle of DNA encoding two key genes into these T cells. One endowed the cells with the ability to home in on the cancer cells. The other encoded a gene from a herpes simplex virus called thymide kinase, or HSV1-tk. The product of the HSV1-tk gene traps a radioactively labeled imaging molecule that can be visualized on a PET scan. Any imaging molecule that is not trapped in the modified T cells is eliminated from the body. A clinical PET-CT scanner tracks the locations of the imaging molecule and therefore the modified T cells.
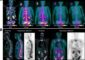
The researchers then returned the modified T cells to the site of the patient's brain tumor over a period of five weeks. The patient received the imaging agent three days after the last infusion of cells. As the researchers had hoped, the subsequent PET-CT scan showed that the T cells had homed in on the tumor. However, they also migrated through the patient's brain to highlight a second, previously unsuspected tumor site. Although this study did not assess the ability of the T cells to kill the tumor cells, the imaging results suggested they at least got to their targets.
“The cells were actually good at finding the tumor,” said Gambhir, who pointed out that the same technique could be used to follow other immune cells or eventually stem cells throughout the body. He plans to collaborate with other researchers at Stanford and elsewhere to not only continue his study with T cells and other tumor types, but also to investigate the movement of therapeutic cells in patients with arthritis and diabetes.
The study could not have been done without the concerted efforts of researchers at Stanford, UCLA, City of Hope and the Food and Drug Administration, Gambhir emphasized. Genetically modifying cells for re-infusion into a human patient requires rigorous quality-control measures and extensive ethical review. The researchers selected the glioblastoma patient for their first attempt because this cell-based therapy trial was already approved by the FDA. Gambhir also had FDA approval on the PET imaging agent.
“It took all of the institutions to come together to make such a complex trial work,” said Gambhir, “but, because we're not limited to just one cell population, the results are tremendously exciting.”
Media Contact
More Information:
http://www.stanford.eduAll latest news from the category: Medical Engineering
The development of medical equipment, products and technical procedures is characterized by high research and development costs in a variety of fields related to the study of human medicine.
innovations-report provides informative and stimulating reports and articles on topics ranging from imaging processes, cell and tissue techniques, optical techniques, implants, orthopedic aids, clinical and medical office equipment, dialysis systems and x-ray/radiation monitoring devices to endoscopy, ultrasound, surgical techniques, and dental materials.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…