What water looks like to DNA

These are snapshots from numerical relaxation of the two-plate system. A red region indicates the solute region without solvent.<br><br>Credit: Reproduced from the Journal of Chemical Physics. See: http://dx.doi.org/10.1063/1.4812839<br>
A team of biochemists and mathematicians have developed a sophisticated geometric model to predict how a biological molecule will interact with water molecules, computing the results up to 20 times faster than other existing approaches.
This new approach may help researchers find new drugs to treat human diseases, said the team, who described their theoretical approach in the Journal of Chemical Physics, which is produced by AIP Publishing.
“Our research explores how water can change the shape of a molecule, how different molecules can get along well in water and, ultimately, how drug molecules can hit targets with the help of water,” says Bo Li, professor of mathematics and senior scientist, National Science Foundation Center for Theoretical Biological Physics, University of California, San Diego.
Biological molecules such as DNA and proteins are the building blocks of living systems, and each molecule consists of many atoms. “How these molecules self-organize is crucial to maintaining a healthy system, because a missing or deformed atom within a molecule can lead to disease,” explained Li.
The human body contains numerous biological molecules, many of which are surrounded by water, which can help change their shape and affect how they interact with other molecules in the body. Up to 60 percent of the human body is water, so it's essential that this solvent be considered.
“Many biological molecules are hydrophobic (water repelling), just like a drop of oil in water, but when mixed they will eventually blend together,” said Li.
Being able to quickly predict the structure of biological molecules in water by using this new theoretical approach should help improve the ability of researchers to identify new targets and may reduce the need for expensive screening of millions of drug molecules in labs.
This work is part of a joint research program initiated in the lab of J. Andrew McCammon, Joseph E. Mayer Professor of Theoretical Chemistry, Distinguished Professor of Pharmacology, and Howard Hughes Medical Institute (HHMI), University of California, San Diego, and has been supported by a grant from the National Institutes of Health and HHMI.
The article, “Phase-Field Approach to Implicit Solvation of Biomolecules with Coulomb-Field Approximation,” authored by Yanxiang Zhao, Yuen-Yick Kwan, Jianwei Che, Bo Li, and J.A. McCammon, is published in the Journal of Chemical Physics. See: http://dx.doi.org/10.1063/1.4812839
ABOUT THE JOURNAL
The Journal of Chemical Physics publishes concise and definitive reports of significant research in the methods and applications of chemical physics. See: http://jcp.aip.org
Media Contact
More Information:
http://www.aip.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
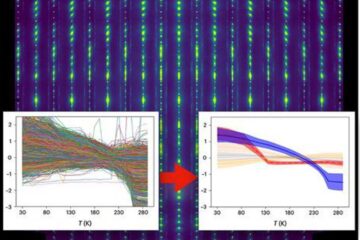
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
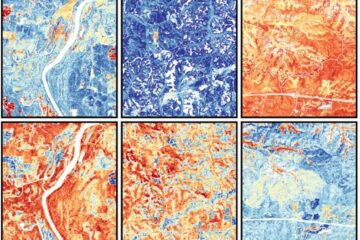

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















