UMD researchers develop tool to better visualize, analyze human genomic data

Next-generation sequencing has revolutionized functional genomics. These techniques are key to understanding the molecular mechanisms underlying cell function in healthy and diseased individuals and the development of diseases like cancer. Data from multiple experiments need to be integrated, but the growing number of data sets makes a thorough comparison and analysis of results challenging.
To visualize and browse entire genomes, graphical interfaces that display information from a database of genomic data—called “genome browsers”—were created. Epiviz offers a major advantage over browsers currently available: Epiviz seamlessly integrates with the open-source Bioconductor analysis software widely used by genomic scientists, through its Epivizr Bioconductor package.
“Prior tools limited visualization to presentation and dissemination, rather than a hybrid tool integrating interactive visualization with algorithmic analysis,” says Héctor Corrada Bravo, assistant professor in computer science at UMD. He also has an appointment in the Center for Bioinformatics and Computational Biology of the university's Institute for Advanced Computer Studies.
Because Epiviz is based on the Bioconductor infrastructure, the tool supports many popular next-generation sequencing techniques, such as ChIP-seq, which is used to analyze protein interactions with DNA; RNA-seq, which reveals a comprehensive snapshot of the abundance of RNAs in cells; and DNA methylation analyses.
Epiviz implements multiple visualization methods for location-based data (such as genomic regions of interest) and feature-based data (such as gene expression), using interactive data visualization techniques not available in web-based genome browsers. For example, because display objects are mapped directly to data elements, Epiviz links data across different visualizations giving users visual insights of the spatial relationships of multiple data sets. The tool is designed to allow biomedical scientists to easily incorporate their own visualizations.
In the Nature Methods paper, Corrada Bravo, UMD computer science doctoral student Florin Chelaru, and undergraduate research assistants from Williams College in Mass. and Washington University in St. Louis used Epiviz to visualize and analyze DNA methylation and gene expression data in colon cancer. Changes in DNA methylation patterns compared with normal tissue have been associated with a large number of human malignancies.
Using Epiviz and Bioconductor, the research team found consistent regions of DNA methylation changes in colon cancer samples generated by the public Cancer Genome Atlas project and similar gene expression in these regions of DNA methylation changes in other cancer types. The results were in agreement with previous experiments, which were conducted by researchers at Johns Hopkins University in collaboration with Corrada Bravo, showing DNA methylation changes across large regions in the colon cancer genome.
“Epiviz helps biomedical scientists meet the challenge of visualizing large genomic data sets while supporting creative data analysis in a collaborative environment,” says Corrada Bravo.
This research was supported by the National Institutes of Health (Award Nos. HG006102 and HG005220), Illumina Corp. and Genentech. The content of this article does not necessarily reflect the views of these organizations.
Héctor Corrada Bravo website: http://www.cbcb.umd.edu/~hcorrada/
The research paper, “Epiviz: interactive visual analytics for functional genomics data,” Florin Chelaru, Llewellyn Smith, Naomi Goldstein and Héctor Corrada Bravo, was published online Aug. 3, 2014 in Nature Methods. http://dx.doi.org/10.1038/nmeth.3038
Media Relations Contact: Abby Robinson, 301-405-5845, abbyr@umd.edu
Writer:
Melissa Brachfeld
University of Maryland
College of Computer, Mathematical, and Natural Sciences
2300 Symons Hall
College Park, Md. 20742
http://www.cmns.umd.edu
About the College of Computer, Mathematical, and Natural Sciences
The College of Computer, Mathematical, and Natural Sciences at the University of Maryland educates more than 7,000 future scientific leaders in its undergraduate and graduate programs each year. The college's 10 departments and more than a dozen interdisciplinary research centers foster scientific discovery with annual sponsored research funding exceeding $150 million.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
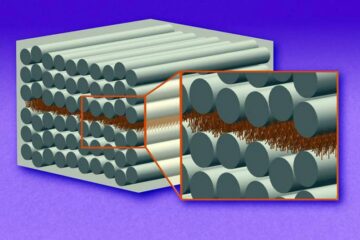

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















