When siblings grow apart

The genomes of higher organisms generally contain numerous genes originating from duplication events. In many cases, the resulting gene pairs maintain essentially parallel functions over the course of evolution, as demonstrated recently by Kousuke Hanada and colleagues from the RIKEN Plant Science Center in Yokohama.
Working with thale cress (Arabidopsis thaliana), the investigators found that evolution often tends to select for duplications that build redundancy into the genome, shielding organisms from potentially disastrous effects of function-altering mutations1. However, this doesn’t tell the full story about gene duplication.
“Knocking out either of two duplicate genes sometimes induces totally different phenotypes, indicating that the two copies have different functions,” says Hanada. This speaks to a process of ‘functionalization’, in which the two duplicate genes either evolve distinct functional profiles or else divide up the functions of the original, pre-duplication gene. Hanada and colleagues have now subjected A. thaliana to further analysis in order to better understand the molecular and evolutionary basis of this ‘morphological diversification’2.
Gene function can be altered either through changes to the encoded protein sequence or modifications to their expression behavior. The researchers began by assessing these characteristics in 398 gene pairs that had undergone functionalization relative to 94 pairs that had not, using sequence and expression data from the published literature and publicly available gene expression databases.
As expected, sequence and expression variability were both found to be significantly higher within gene pairs that had undergone some degree of functionalization. However, there was also a striking difference in the relative contribution of these factors to morphological diversification. “Our analysis suggested that changes [in] expression pattern play the minor role—between 33 and 41%—and that changes [in] protein sequence play the major role—between 59 and 67%,” says Hanada. “This result is most surprising; most people believed that changes in expression pattern play the major role because such changes are essential for development.”
The investigators are now keen to apply their classification strategy to other organisms, including the fruit fly and mouse, in an effort to determine whether similar evolutionary patterns exist. “I expect that changes of expression pattern are more important in complex organisms than simple organisms, but I do not know the real answer yet,” says Hanada. Collectively, the resulting data could inform development of tools that enable scientists to better understand the evolution of gene function based on observed sequence and expression changes.
The corresponding author for this highlight is based at the Gene Discovery Research Group, RIKEN Plant Science Center
1. Hanada, K., Kuromori, T., Myouga, F., Toyoda, T., Li, W.-H. & Shinozaki, K. Evolutionary persistence of functional compensation by duplicate genes in Arabidopsis. Genome Biology and Evolution 409, 409–414 (2009).
2. Hanada, K., Kuromori, T., Myouga, F., Toyoda, T. & Shinozaki, K. Increased expression and protein divergence in duplicate genes is associated with morphological diversification. PLoS Genetics 5, e1000781 (2009)
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
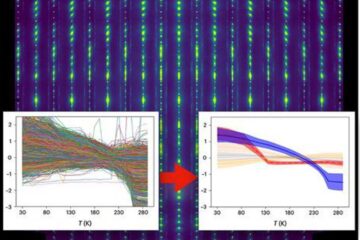
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
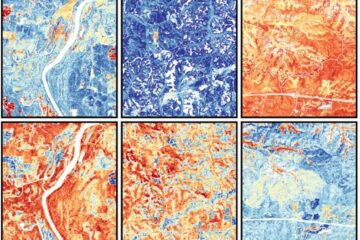

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















