New role for phosphorylation in heterochromatin
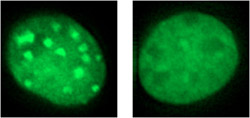
HP1á accumulates in heterochromatic regions in wildtype cells (left), but not in cells engineered to lack phosphorylation of the protein’s N-terminal region (right).<br>
A great many cellular processes are switched on or off by the modification of a given enzyme or other protein by addition of a phosphate molecule, known as phosphorylation. This regulatory activity occurs widely in the cytoplasm, but can take place in the nucleus as well. Recent work has shown the HP1á, a protein that guides the formation of heterochromatin, a form of the DNA-protein structure know as chromatin, is also subject to this post-translational modification, but the biological meaning of this event has remained unresolved.
A new study by Kyoko Hiragami-Hamada and colleagues in the Laboratory for Chromatin Dynamics (Jun-ichi Nakayama, Team Leader), working in collaboration with labs in Kobe University, Kwansei Gakuin University, and AIST, have shown that phosphorylation of HP1á boosts its ability to bind to heterochromatin, resulting in stabilization of chromosomes. Published in Molecular and Cellular Biology, this work opens up new insights into the interplay between protein modification and chromatin dynamics.
HP1 was first identified in Drosophila, and is now recognized as a highly conserved regulator of transcriptional repression in heterochromatin in eukaryotes. The protein has two similar binding domains: a chromodomain (CD) region in its N-terminal region, and a chromo shadow domain (CSD) in the carboxyl end. The CD binds to a methylated site on histone H3 (H3K9me3), and importantly the CSD promotes binding to other HP1 proteins, which is essential to the formation and maintenance of heterochromatin. It has been suspected that HP1 may also be regulated by phosphorylation as well, but this has never clearly been shown.
Hiragami-Hamada began by testing for binding activity between mammalian HP1 variants HP1á, HP1â and HP1ã and H3K9me3, and found that while phosphorylation had no effect on the strength of this association in HP1â or HP1ã, binding of HP1á to methylated histone H3 appeared to be phosphorylation-dependent. She next used an electrophoresis technique known as Phos-tag-PAGE to examine HP1á in its phosphorylated state, and found not only that multiple sites on the protein were thus modified, but that phosphorylation increased during cell division as well.
To resolve the specific phosphorylation sites, she next used Phos-tag-PAGE in combination with targeted amino acid substitutions and identified serine 14 in the N-terminal and serine 93 in the protein’s hinge region as phospho-targets. Mass spectrometry revealed additional sites in serine 11 – 13 in the N-terminal as well. Tests of binding affinity showed that it was the N-terminal sites, but not S93 in the hinge, that are responsible for binding to H3K9me3 in a phosphorylation-dependent manner. Interestingly, while this modification was important for establishing the interaction, it was dispensable in its maintenance.
With a better understanding of the biochemistry behind this interaction, Hiragami-Hamada next turned to phosphorylated HP1á’s role within the cell. She found that, when phosphorylation of S14 was prevented by an amino acid substitution, HP1á accumulation at heterochromatic regions was reduced. Similar experiments on N-terminal serines 11 to 13 showed that their phosphorylation also plays a role in targeting HP1á to heterochromatin. When the team replaced all four serines 11 through 14 with substitute amino-acids in cultured mouse cells, they observed an increase in chromosomal abnormalities. Serine 93 in the hinge region in contrast had no such effects.
“This work suggests that the chromodomain on its own is not sufficient for binding the methylated histone; the connections needs to be strengthened by phosphorylation of sites on the HP1á N-terminus, which interestingly contributes to chromosomal stabilization as well” says Nakayama. “We’re now curious about how this phosphorylation is regulated, and what role, if any, it plays in serine 93.”
[ Contact ]
Douglas Sipp : sipp(at)cdb.riken.jp
TEL : +81-78-306-3043
RIKEN CDB, Office for Science Communications and International Affairs
Journal information
Mol Cell Biol. 2011 Mar;31(6):1186-200. Epub 2011 Jan 18.
'N-Terminal Phosphorylation of HP1{alpha} Promotes Its Chromatin Binding.'
Hiragami-Hamada K, Shinmyozu K, Hamada D, Tatsu Y, Uegaki K, Fujiwara S, Nakayama J.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Combatting disruptive ‘noise’ in quantum communication
In a significant milestone for quantum communication technology, an experiment has demonstrated how networks can be leveraged to combat disruptive ‘noise’ in quantum communications. The international effort led by researchers…

Stretchable quantum dot display
Intrinsically stretchable quantum dot-based light-emitting diodes achieved record-breaking performance. A team of South Korean scientists led by Professor KIM Dae-Hyeong of the Center for Nanoparticle Research within the Institute for…

Internet can achieve quantum speed with light saved as sound
Researchers at the University of Copenhagen’s Niels Bohr Institute have developed a new way to create quantum memory: A small drum can store data sent with light in its sonic…





















