Mutating the entire genome

Genes account for only 2.5 percent of DNA in the human genetic blueprint, yet diseases can result not only from mutant genes, but from mutations of other DNA that controls genes. University of Utah researchers report in the journal Nature Genetics that they have developed a faster, less expensive technique for mutating those large, non-gene stretches of DNA.
“Diseases are known to occur as a consequence of deleting non-gene DNA sequences, and this new method allows us to evaluate what these sequences do,” says Mario Capecchi, distinguished professor and co-chair of human genetics at the University of Utah and an investigator for the Howard Hughes Medical Institute (HHMI).
The new method “is significant because it makes it practical to do this for a vast amount of the total genome,” he adds.
Because mice are used to study human disease, “we want to know the function of every piece of DNA in the mouse genome,” says Sen Wu, a postdoctoral fellow in human genetics at the University of Utah and HHMI. “The best way to know the function of the genetic blueprint is by removing part of the DNA and seeing what goes wrong. We have found a way to do this job on a large scale that is simple and practical.”
Capecchi says: “We developed a method for deleting any piece of DNA from the mouse genetic blueprint very efficiently.”
The new method for mutating large, non-gene stretches of DNA is outlined in this week’s online edition of Nature Genetics. Capecchi and Wu conducted the research with two other University of Utah human geneticists: Guoxin Ying, a postdoctoral fellow, and Qiang Wu, an assistant professor (and no relation to Sen Wu).
In the journal paper, the University of Utah scientists report:
They found a way to delete or duplicate moderately long to very long pieces of DNA and make those mutations happen much more frequently than other methods can. That makes it easier to find out what defects or diseases arise due to such mutations, and thus what the DNA does normally.
They devised a much more efficient method for mixing and recombining pieces of two chromosomes, making it easier to breed mice with human cancers. Such mice are needed to develop new treatments.
Vast, Non-Gene Stretches of the Genetic Blueprint
The genome is the genetic blueprint of a living organism. It is made of deoxyribonucleic acid, or DNA, a twisting, double ladder-shaped molecule made from numerous “base pairs” of four building blocks: nucleic acids designated A, C, G and T.
DNA in each human cell has about 3 billion base pairs, arranged into 23 pairs of chromosomes. Those chromosomes include roughly 20,000 genes, which carry the code needed for cells to produce the proteins used to make body parts and carry out most functions in living organisms. Genes typically have 50,000 base pairs but can range in size from a few thousand to 300,000 base pairs, Capecchi says.
The genes include only about 2.5 percent to 3 percent of human DNA. In between the genes are vast stretches of “non-coding” or non-gene DNA.
Some pieces of non-gene DNA are “regulatory sequences” that turn genes on or off, or turn their activity up or down like a dimmer switch. Other DNA sequences are responsible for folding and packing DNA into the nucleus of each cell.
Five percent to 50 percent of DNA is considered useless junk, “depending on who you ask, although probably at least half of the ‘junk’ DNA is going to end up having a pretty important function,” Capecchi says.
Capecchi is known for developing gene targeting, in which mice are bred with a gene disabled or “knocked out” to see what goes wrong, thus revealing the gene’s normal function and how disease makes it malfunction. But DNA regulatory sequences between genes also can mutate to cause disease by making the genes they control malfunction.
“We know how to knock out genes, and that technology is very good at changing any given gene in a designed manner,” Capecchi says. “The negative side is, it takes work and money to do that.” The new method is “a cheaper way of knocking out a gene as well as regulatory sequences.”
Capecchi says it now costs about $10,000 to genetically engineer a mouse with a particular gene or other DNA sequence knocked out. So using existing technology to knock out the estimated 20,000 mouse genes would cost $200 million, and knocking out the roughly 300,000 non-gene DNA sequences in mice would cost $3 billion, he adds.
The new method is a faster, cheaper way of mutating DNA, costing $200 to create each mouse with a mutant gene or other DNA sequence, although the savings are not proportionate because more mutant mice must be bred to obtain desired mutants, he says.
Capecchi says the new technique could speed a National Institutes of Health effort to mutate all mouse genes in cell culture and create 900 new lines of mutant mice by 2010 – an effort NIH says “will be extremely useful for the study of human disease.”
Simpler, Faster Mutations in the Whole Mouse Genome
Existing gene-targeting methods for deleting large segments of DNA are time-consuming and expensive because they require researchers to make multiple manipulations in mouse embryonic stem cells, which means those cells are less likely to yield mice with the desired stretch of DNA disabled, Capecchi says.
To quickly, cheaply mutate genes or none-gene DNA, Wu and Capecchi used short pieces of DNA known as loxP. Pieces of loxP act like signposts saying, “Cut the DNA between these two signposts.” In the new method, loxP is inserted in a chromosome on one mouse, and in a different position on the same chromosome in a second mouse, which also has a gene named Cre in its cells. When the mice are bred, an offspring mouse has loxP DNA on two sites on the same chromosome and also has the Cre gene. The protein made by Cre acts like a knife, cutting the DNA wherever loxP is found.
The offspring mouse, in turn, is bred with a normal mouse. In a “remarkably high” average of 10 percent of the offspring, the desired large stretch of DNA is either deleted or duplicated so scientists can see what goes wrong in the mutant mouse and thus learn the normal job of that piece of DNA.
Capecchi says the same method also can be used to make two chromosomes break and recombine, with each new chromosome containing a piece of each of the two old ones. The process, “translocation,” also can create new genes from two other genes. Many cancers – certain sarcomas, leukemias and lymphomas – start with translocation.
Existing methods result in the desired translocation of two chromosomes in one of every 1 million to 10 million cells cultured, Capecchi says, while the new method produces the desired result in one of every 100 cells. So it is 10,000-100,000 times better.
That is important, because multiple molecular steps are needed for a mouse to develop a human cancer – and such mice are used to learn how to treat or prevent cancers, says Capecchi. In many cancers, translocation of two chromosomes is the first step, and if it occurs only in one of every million cells, not enough cells are present in which subsequent steps can occur so that cancer develops.
“If we now improve that 10,000-fold, the pool of cells [with translocated chromosomes] may be large enough to create a mouse that develops the cancer,” so the new method makes it easier to breed mice with human cancers, Capecchi says.
‘Jumping Genes’ Help Make More Mutant DNA
Wu and Capecchi improved their new method by using it in combination with a “transposon” or so-called “jumping gene” named piggyBac, which comes from a moth. They used piggyBac to insert pieces of loxP DNA randomly into numerous spots on the mouse genome. So scientists easily can breed multiple generations of mice with loxP surrounding various large stretches of DNA.
Then the first part of the new method is used: The mice are bred with mice that carry the Cre gene so any large stretch of DNA that is located between two short pieces of loxP DNA can be mutated.
Because the whole mouse genome has been sequenced – the order and location of base pairs determined in detail – scientists can identify locations in individual mice where the jumping genes have carried pieces of loxP DNA. Then, they can select a mouse with loxP on both sides of the large piece of DNA they want to mutate and study.
If the jumping gene carrying loxP hops into the middle of a gene, then the gene will be mutated. So while the method began as a way to delete large stretches of non-gene DNA, it has the bonus of also being able to disable genes more easily and inexpensively than existing knockout techniques, Wu says.
In their study, Wu and Capecchi demonstrated that by using a jumping gene to mutate a mouse gene involved in bone formation, resulting in mutant mice with abnormally short limbs and tail (see photo with this news release).
Media Contact
More Information:
http://www.utah.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
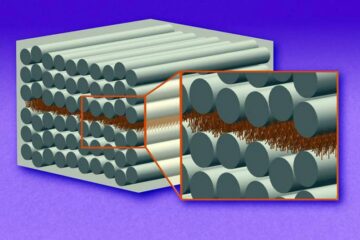

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
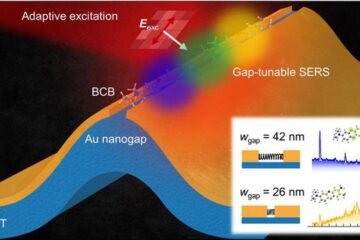
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















