Researchers discover mutations in the progranulin gene cause frontotemporal dementia

Progranulin is a type of protein known as a growth factor. Production of too much progranulin has been associated with cancer. So the gene that codes for progranulin was not an obvious one to sequence in order to look for mutations that cause neurodegenerative disease. However, researchers solved a ten-year genetic puzzle when they found mutations in the gene explain a large number of FTD cases in North America and Europe.
Although researchers found the age of onset in people carrying one of these mutations can come as early as their 50s or as late as their 90s, it is almost certain that anyone with an identified progranulin gene mutation will develop FTD at some point in life if they live long enough.
FTD, the second most common form of dementia after Alzheimer's disease, is a group of brain disorders that affect the frontal and temporal lobes of the brain, which control personality and speech. One or both of these functions may be affected. Patients may exhibit apathetic or uninhibited behavior and increasing lack of self-awareness. Patients may also lose the ability to put words together to form intelligible sentences. Speech decreases, and patients may become mute. However, patients usually retain memory until later in the disease course. This differentiates FTD from Alzheimer's disease, where memory function is affected early on.
In 1996 researchers first linked a genetic cause for FTD to chromosome 17. In 1998 Mayo Clinic neurobiologist, Michael Hutton, Ph.D., and others discovered mutations in a gene on chromosome 17 that codes for a protein called tau. Investigators discovered mutations in this gene cause the disease in patients from a number of families with a history of FTD. However, many FTD-affected families with genetic linkage to chromosome 17 lacked these mutations, so researchers continued to hunt for an additional culprit gene or genes. “It was like looking for two needles in the same haystack, and you didn't know you were looking for the second one until you found the first one,” Hutton says.
Hutton led a group of collaborators within Mayo Clinic, the University of British Columbia and Vancouver Coastal Health Research Institute in Vancouver, Canada, and the University of Manchester in the United Kingdom. They analyzed over 80 genes close to the tau gene in FTD-affected and unaffected individuals from a large Canadian family with linkage to chromosome 17, but they failed to find any disease-causing mutations. However, when they sequenced the progranulin gene, located in the same genetic region, they found the first mutation. Subsequent analysis of 42 more FTD families identified a total of nine different mutations in the progranulin gene. All of the mutations effectively knock out one copy of the gene, and therefore its ability to direct production of progranulin. (Gene sequencing is the process of determining the order of the four DNA bases. Each of the approximately 30,000 human genes has its own unique order.)
“What we've found is a little bit different than what we've found in other common neurodegenerative diseases,” Hutton says. “What we're looking at here is simply the loss of progranulin that is causing the disease.” Other neurodegenerative diseases, such as Alzheimer's disease, Parkinson's disease and even FTD caused by mutations in the tau gene, are characterized by the accumulation of disease-specific proteins within surviving brain cells. “Here it's the other way around,” Hutton says. “One copy of the progranulin gene has been knocked out by the mutation, and therefore we have less progranulin produced, which is enough on its own to cause the disease.”
The mutations not only reveal the mechanism that causes the disease, they point to potential for a cure. “Replacing progranulin is the obvious therapeutic approach,” Hutton says. That might be possible through gene therapy. Or by understanding the process that regulates progranulin expression, researchers may find ways to increase progranulin production from the surviving copy of the gene.
There are many growth factors required for neuronal function, and although Hutton and his colleagues don't yet know what role progranulin plays in the normal function of these brain cells, he says this discovery implies there may be other brain disorders, such as Lou Gehrig's disease (amyotrophic lateral sclerosis) in which the loss of certain growth factor-type proteins can actually give rise to the disease.
Because of its apparent role in neuronal function, Hutton's lab has begun to investigate whether normal variability in the progranulin gene influences the risk of developing Alzheimer's disease or Parkinson's disease. He asks, “If you have a particular, common variant in the progranulin gene, does that mean you are protected from getting Alzheimer's disease or a lower risk, because your neurons are better able to withstand the kind of damage they get from accumulation of amyloid beta?” (The amyloid beta protein is the principal component of the senile plaques that develop in brain cells of people with Alzheimer's disease.)
Authors contributing to the paper to be published by Nature are: Matt Baker, Jennifer Gass, Rosa Rademakers, Jennifer Adamson, Ashley Cannon, Stacey Melquist, Dennis Dickson, Zdenek Berger, Jason Eriksen, Todd Robinson, Cynthia Zehr, Chad A. Dickey, Richard Crook, Eileen McGowan, Mike Hutton, Department of Neurosciences, Mayo Clinic College of Medicine; Bradley Boeve, Department of Neurology, Mayo Clinic College of Medicine; Ian R. Mackenzie, Department of Pathology, University of British Columbia; Caroline Lindholm, A. Dessa Sadovnick, Howard Feldman, Division of Neurology, University of British Columbia; Emily Dwosh, Department of Medical Genetics, University of British Columbia; Stuart M. Pickering-Brown, Sara Rollinson, Division of Laboratory and Regenerative Medicine, Department of Medicine, University of Manchester; Julie Snowden, Anna Richardson, David Neary, David Mann, Centre for Clinical Neurosciences, University of Manchester.
A second paper, from researchers at the University of Antwerp in Belgium, describing similar findings, will be published at the same time in Nature.
This research was funded by the National Institute on Aging, Mayo Foundation and the Robert H. and Clarice Smith Fellows program.
Media Contact
More Information:
http://www.mayo.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
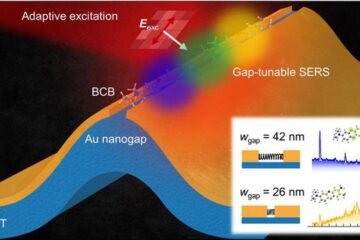
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















