Study finds 60 new genes controlled by DNA snippet

May lead to future treatments for heart failure, nerve disorders
Researchers worldwide are seeking to define ancient sections of our genetic code that may soon be as important to medical science as genes. A new wave of research is concerned with, not how genes work, but how small regulatory DNA sequences tell genes where, when and to what degree to “turn on.” As part of this effort, researchers at the University of Rochester Medical Center scanned through the vast human DNA code to reveal for the first time 60 genes influenced by one such sequence, according to an article published today in the journal Genome Research.
Growing knowledge of how regulatory sequences control gene behavior has the potential to create new classes of treatment for nerve disorders and heart failure. Such sequences may also help to explain why humans are so complex, despite having one-fifth as much genetic material as wheat for instance. Medical center researchers are working on just one of more than 100 regulatory sequences identified so far, each the subject of intense study.
“Most people don’t realize that genes make up a very small percentage of the human DNA code,” said Joseph M. Miano, Ph.D., senior author of the journal paper and associate professor within the Cardiovascular Research Institute at the medical center. “Genes are relatively straightforward compared to what lies ahead. We believe that the real genetic gymnastics, the real intelligence of our system, is controlled by tiny bits of genetic material that tell genes what to do.”
“Junk DNA” No More
Genes are the chains of deoxyribonucleic acids (DNA) that encode instructions for the building of proteins, the workhorses that make up the body’s organs and carry its signals. The Human Genome Project, which first reported results in 2001, produced a near complete listing of the DNA sequences that make up all human genes (the genome). Key project findings included that human genetic material consists of about 3 billion base pairs, the “letters” that make up the genetic code. Researchers also concluded that genes, specific batches of code that direct protein construction, comprise just about 2 percent of all human DNA. A central question in genetics has become: what does the remaining 98 percent of human genetic material do?
Regulatory sequences are emerging as an important part of the non-gene majority of human genetic material, once thought of as “junk DNA.” A new frontier in genetic research is the defining of the regulome, the complete set of DNA sequences that regulate the behavior of genes. DNA segments that code for proteins average 200 base pairs in length, whereas regulatory sequences typically include just six to 10 base pairs, making them hard to find.
As a human embryo develops from a single cell into tens of billions of cells, DNA must be read and copied again and again to supply each cell with its needed copy. Over time, random changes, or mutations, are inserted into the code during the copying process. Some mutations bring survival advantages and others cause disease. Most known genetic diseases identified to date result from a mutation within a gene that directs protein construction, but that may soon change.
“We believe more and more disease-causing mutations will be found within regulatory sequences that control genes turning on or off,” Miano said. “We therefore are very interested in defining as many functional regulatory elements as we can to help geneticists pinpoint a growing number of disease-causing mutations.”
Study Details
In Miano’s study, the regulatory sequence under examination was the CArG box. The nucleotide building blocks of DNA chains may contain any one of four nucleobases: adenine (A), thymine (T), guanine (G) and cytosine (C). Any sequence of code starting with 2 Cs, followed by any combination of 6 As or Ts, and ending in 2 Gs is a CArG box.
According to Miano, there are 1,216 variations of CArG box that together occur approximately three million times throughout the human DNA blueprint.
CArG boxes exert their influence over genes because they are “shaped” to partner with a nuclear factor called serum response factor (SRF) and several other proteins within a genetic regulatory network. Throughout a human life, such networks are believed to “decide” the timing and location of all gene expression, the process through which genetic information is converted into templates for protein construction.
The current study, funded through a grant from the National Heart, Lung and Blood Institute, sought to survey the human and mouse genome databases created by the Human Genome Project to find all CArG boxes that regulate genes. The sheer amount of information involved requires that such studies use high-powered computer programs to screen data. In this case, researchers used a high-speed screening to expand the definition of the functional mammalian CarGome, the complete set of CArG boxes that regulate genes.
In collaboration with Christian Stoeckert, Ph.D., associate professor of Genetics at the University of Pennsylvania, Miano’s team designed a set of criteria that a given piece of DNA had to meet in order to be considered a functional CArG box. Thanks to their work and that of several other labs, they knew going in all the CArG box variations and how close they typically lie to the genes they regulate (within 4,000 base pairs).
The data-screening tool also employed comparative genomics, the study of relationships between the DNA of different species. When a piece of genetic material, gene or regulatory segment, is conserved by evolution from mice to humans it suggests that the segment has a valuable function. Miano’s screen required that CArG boxes shared by humans and mice be included in his expanded version of the CarGome. CArG boxes identified by the computer screen were then tested to see if they indeed interacted with SRF and changed the behavior of genes as predicted.
This approach resulted in the disclosure of more than 100 hypothetical CArG boxes and the same number of genes previously unknown to be targeted by CArG-SRF. Of those, 60 CArG boxes have been validated as exerting influence over a gene. Adding the newly confirmed segments to those already known, authors of the study now define the functional mammalian CarGome as 161 sequences, a 55 percent increase from the old definition.
Of the genes newly found to be regulated by CArG-SRF, more than half encode for cytoskeletal or contractile proteins. Past studies have shown that CArG-SRF network is vital to the development of the cellular “skeletons” that maintain cell shape and enable cell motion. Being present in nearly every cell and throughout the human genome, the CArG-SRF system is believed to contribute to disease in many bodily tissues.
In cardiology, studies show that a lack of SRF-CArG causes cardiomyopathy, a weakening of heart muscle cells’ ability to contract. That in turn reduces the pumping strength of heart muscle and leads to heart failure, according to recent studies. Can cardiomyopathy be reversed by manipulating CArG-SRF? CArG sequences also appear near genes that direct the building of nerve cells and blood vessels, suggesting they may be involved in diseases affecting those tissues as well.
In the larger picture, regulatory sequences may help to explain why humans have just 25,000 genes when, given the degree of human complexity, researchers had expected to find more than 100,000. Regulatory sequences may be part of the answer because they enable a single gene to produce the same protein at different times, places and concentrations with subtly different roles.
“Humans share about one quarter of their genes with fish,” Miano said. “Something must be at work to explain why we are so many times more complex. Regulatory sequences offer one of several emerging explanations for how we do more with fewer genes.”
Media Contact
More Information:
http://www.urmc.rochester.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
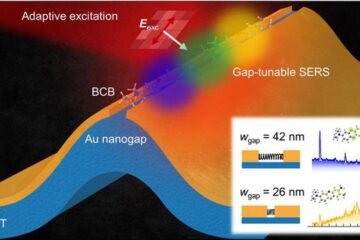
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















