How E. coli bacterium generates simplicity from complexity
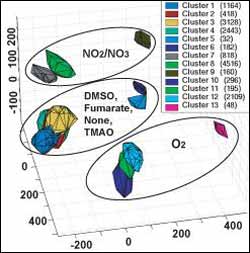
This statistical projection of E. coli’s computation-based activity profiles permits researchers to visualize the "space" of transcriptional regulation of genes involved in metablism and metabolic regulation. The clusters’ positions are a function of the available electron acceptor, indicated by the ellipses, the carbon "food" source, and to a lesser degree by the source of nitrogen. (The number in parenthesis by each of the 13 clusters is the numbers of different activity profiles in the cluster.)
The ubiquitous and usually harmless E. coli bacterium, which has one-seventh the number of genes as a human, has more than 1,000 of them involved in metabolism and metabolic regulation. Activation of random combinations of these genes would theoretically be capable of generating a huge variety of internal states; however, researchers at UCSD will report in the Dec. 27 issue of Proceedings of the National Academy of Sciences (PNAS) that Escherichia coli doesn’t gamble with its metabolism. In a surprise about E. coli that may offer clues about how human cells operate, the PNAS paper reports that only a handful of dominant metabolic states are found in E. coli when it is “grown” in 15,580 different environments in computer simulations.
“When it comes to genomes, a great deal of complexity boils down to just a few simple themes,” said Bernhard Palsson, a professor of bioengineering at UCSD’s Jacobs School of Engineering and co-author of the study, which was made available online Dec. 15. “Researchers have confirmed the complexity of individual parts of biochemical networks in E. coli and other model organisms, but our large-scale reconstruction of regulatory and metabolic networks involving hundreds of these parts has shown that all this genetic complexity yields surprisingly few physiological functions. This is possibly a general principal in many, if not all, species.”
This statistical projection of E. coli’s computation-based activity profiles permits researchers to visualize the “space” of transcriptional regulation of genes involved in metablism and metabolic regulation. The clusters’ positions are a function of the available electron acceptor, indicated by the ellipses, the carbon “food” source, and to a lesser degree by the source of nitrogen. (The number in parenthesis by each of the 13 clusters is the numbers of different activity profiles in the cluster.)
Palsson and his colleagues at UCSD, postdoctoral fellows Christian L. Barrett and Christopher D. Herring, and Ph.D. candidate Jennifer L. Reed, created a computer model of an E. coli cell based on the experimental results of thousands of previous experiments, some of which were completed decades ago. “The goal of this study was to comprehensively simulate all the possible molecular interactions in a well studied strain of E. coli to gain a global view of the range of functional network states,” said Barrett. “Complex cellular networks can potentially generate lots of different behaviors, but we find that cells utilize only a few of them.”
Barrett, Palsson, Herring, and Reed simulated the behavior of 1,010 of E. coli’s 4,200 genes. This particular subset of the bacterium’s genome is tightly organized into interacting networks involved in metabolism or regulation of gene activation, or transcription. These linked networks are devoted to sensing, ingesting, and degrading potential “food” in the form of sugars and other energy-rich organic molecules.
E. coli must also have an efficient way to eliminate waste products. It, like all living things, generates energy in a process that involves the removal of electrons from food molecules and attaching them to acceptor molecules. For aerobic organisms, the final electron acceptor is usually oxygen, which is converted into water in the process.
E. coli can grow with or without oxygen, using nitrate or other molecules as its final electron acceptor. “We found that the type of terminal electron acceptor in the growth environment and the presence or absence of glucose is very important to E. coli,” said Barrett. “Our simulations show that these two factors are key determinants of how the bacterium organizes itself.”
Barrett, Palsson, and their colleagues simulated the “functional states” of E. coli’s metabolic and transcriptional regulatory networks in the 15,580 environments of food sources and electron acceptors. To their surprise, no matter what carbon source it ingests or electron acceptor used, E. coli exhibits only six distinct functional states.
“This study gives a systems biology view of how a phenotype, or a ‘network state’ advantageous to a microorganism is comprised of a tiny subset of a much larger universe of possibilities as provided for in the genome,” said Palsson. “On a high level we can say that E. coli is obsessed with how it breathes and whether or not glucose is available to eat. All of its genetic complexity basically enables it to generate a nice steady state for itself regardless of what it has to live on.”
Higher organisms have larger genomes and much more complexity, but Palsson noted that several theoretical studies predict that even eukaryotic cells will exhibit a relatively small number of functional states. “When we uncover the regulatory networks in eukaryotes, including human, we will most likely be able to use computer simulations to uncover the different possible cell types in a manner similar to what was done in our work with E. coli,” said Palsson.
Christian L. Barrett, Christopher D. Herring, Jennifer L. Reed, and Bernhard O. Palsson, “The global transcriptional regulatory network for metabolism in Escherichia coli exhibits few dominant functional states” (2005). 102 (52), pp 19103–19108.
Media Contact
More Information:
http://www.ucsd.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…





















