Protein aggregates in Lou Gehrig’s disease linked to neuron death

French neurologist Jean-Martin Charcot first described amyotrophic lateral sclerosis (ALS) in 1869, but, nearly 140 years later, little is known about the cause of the devastating neurodegenerative disease, and there is no cure.
What is known about Lou Gehrig’s disease, as it is commonly called, is that misfolded and damaged proteins clump together in cells to form aggregates and motor neurons die. But scientists have long debated whether or not the protein aggregates actually kill the cells.
Now a research team at Northwestern University, using mammalian neurons and live-cell time-lapse spectroscopy, has become the first to clearly link the presence of the ALS-associated mutant SOD1 protein aggregates with neuronal cell death. This evidence could help explain the disease process and eventually lead to new therapeutics.
In the study, published this month in the Journal of Cell Biology, the scientists looked one at a time at neuronal cells expressing the mutant SOD1 protein and found that in cells where the protein accumulated and aggregates formed, 90 percent of the cells went on to die. (They died between six and 24 hours after aggregates were visually detected.) Cells that did not form aggregates did not die.
The study also provides a new understanding of the structure and composition of the deadly aggregates — one of the first studies to do so.
“We found that these aggregates are quite peculiar and very different from the aggregates formed in Huntington’s disease,” said Richard I. Morimoto, Bill A. and Gayle Cook Professor in Biological Sciences, who led the study. Morimoto is an expert in Huntington’s disease and on the cellular response to damaged proteins.
“In Huntington’s, the aggregate is very dense and impenetrable and binds irreversibly with other molecules in the cell,” he said. “In ALS, the aggregates are amorphous, like a sponge. Other proteins can go through the structure and interact with it, which may help explain why mutant SOD1 is so toxic.” Morimoto believes this surprising finding indicates that the structure of aggregates associated with other neurodegenerative diseases such as Parkinson’s and Alzheimer’s will be found to be different as well.
Looking at individual cells in a population, the researchers also found that cells side by side did different things. In cells expressing the same amount of damaged protein, some cells formed aggregates and died and others did not form aggregates and lived. Only a certain subset of at-risk cells went on to lose function and die.
“It would be terrifying if 100 percent of the cells expressing mutant proteins died,” said Morimoto. “This means that in many cases the cell’s protective machinery suppresses the damaged proteins, keeping the cell healthy. This discovery will be important to scientists looking to develop genetic suppressors and therapeutics.”
Morimoto’s team focused on SOD1 because it is a form of familial (hereditary) ALS in which a mutation in just one gene and its associated protein has devastating consequences to the cell. (Approximately 10 percent of ALS cases are familial.) This provides experimentalists with a powerful framework. For the other 90 percent the disease is not the result of one mutation but rather a series of many genetic events that debilitate motor neurons. With non-familial forms it is extremely difficult to design hypothesis-based experiments, said Morimoto.
The next question the researchers plan to address is what are the events that lead to cell death once mutant SOD1 protein aggregates form in the cell? This knowledge would help scientists identify small molecules that could halt, arrest or reverse the disease process.
In addition to Morimoto, other authors on the Journal of Cell Biology paper are Carina I. Holmberg, Soojin Kim, Gen Matsumoto (lead author) and Aleksandar Stojanovic, all formerly from Northwestern University.
Media Contact
More Information:
http://www.northwestern.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
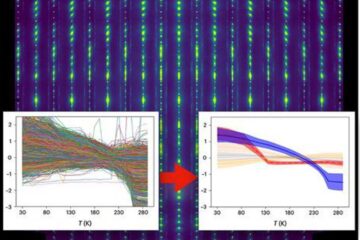
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
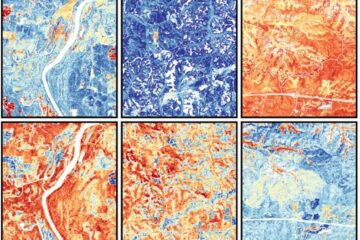

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















