Spiders Help Scientists Discover How Muscles Relax
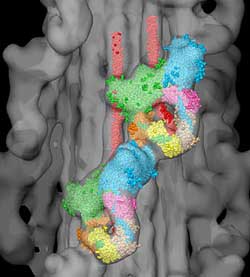
Atomic Model of the Thick Filament from Tarantula Striated Muscle - The surface of the three-dimensional reconstruction of the thick filament is shown in gray, together with the atomic models of two myosin molecules, indicating the intramolecular and intermolecular interactions. Image: John Woodhead
Using muscle tissue from tarantulas, an HHMI international research scholar and his colleagues have figured out the detailed structure and arrangement of the miniature molecular motors that control movement. Their work, which takes advantage of a new technique for visualizing tissues in their natural state, provides new insights into the molecular basis of muscle relaxation, and perhaps its activation too.
“We have solved the structure of the array of miniature motors that form our muscles and found out how they are switched off,” said Raúl Padrón, a HHMI international research scholar in the Department of Structural Biology at the Venezuelan Institute for Scientific Research (Instituto Venezolano de Investigaciones Científicas or IVIC) in Caracas, Venezuela.
The findings are reported in the August 25, 2005, issue of the journal Nature.
Padrón and his colleagues focused their studies on striated muscle—the type of muscle that controls skeletal movement and contractions of the heart. Striated muscles are made of long cylindrical cells called muscle fibers. Within the fibers, millions of units known as sarcomeres give rise to movement of skeletal muscles. Sarcomeres are composed mainly of thick filaments of myosin, the most common protein in muscle cells, responsible for their elastic and contractile properties. The thick filaments are arranged in parallel with thin filaments of another muscle protein, actin. When the actin and myosin filaments slide along one another, the muscle contracts or relaxes.
Padrón’s study focused on the long, rod-shaped myosin of the thick filaments. The heads of these myosin rods project outward from the thick filament to connect with and move actin filaments during contraction of a muscle.
The structural studies were done using tarantula striated muscle, which the team has been studying since the 1980s. Striated muscles from the large, hairy spiders contain filaments that are particularly well ordered, making them easier to study structurally than the more disorganized filaments found in vertebrate striated muscle, Padrón explained.
Padrón and Lorenzo Alamo at IVIC partnered with Roger Craig, John Woodhead and Fa-Qing Zhao at the University of Massachusetts Medical School to use cryo-electron microscopy to answer questions about the thick filament’s structure, questions that could not be answered with existing electron microscopy techniques.
Standard electron microscopy requires dehydration and staining of a tissue sample, which modifies the structure of the specimen and distorts its shape. Cryo-electron microscopy avoids these problems by rapidly freezing the sample. Using the new technique, the researchers were able to visualize the muscle tissue in a form closer to its structure in the body than had previously been possible.
It took several years to refine the techniques required to preserve the thick muscle filaments in their relaxed state. Even then, the researchers faced mathematical difficulties in calculating a three-dimensional map of the filaments. In 2004, using a new approach that Edward Egelman at the University of Virginia Health Sciences Center had developed to create the map, they soon had their structure. “The new reconstruction was very detailed; we were all amazed with the level of detail that it showed,” Padrón said.
The structure provides crucial new details. A twisting, symmetrical arrangement of myosin heads spaced around the filament’s circumference surrounds a backbone made up of 12 parallel strands or sub-filaments. “This is the first time that the structure of the backbone has been clearly seen in any thick filament reconstruction,” the researchers wrote.
“The structure reveals how the helices of the myosin heads are formed and maintained and how the filaments are switched off due to interactions between the myosin heads Padrón explained. “It also opens the way to understanding how the thick filaments are activated.”
The details of their model permitted the researchers to explain how, in relaxed muscle, the heads of each myosin molecule are inhibited from interacting with actin by interacting with each other instead. When muscle is activated, they suggest, the bonds between the myosin heads are broken. This frees each head to interact with actin and cause muscle contraction.
“We have focused on the relaxed muscle to understand the structure of the thick filaments when they are not involved in contraction, but rather fully ordered—a state more amenable to understanding their structure,” Padrón said. “Solving the structure of the relaxed state will allow us to investigate how these filaments are activated when they are switched on.”
The scientists were surprised to find that the atomic structure of isolated myosin molecules from vertebrate smooth muscle—the type of muscle found in the digestive tract, bladder, arteries, and veins—closely matched their invertebrate striated muscle myosin filament. Kenneth Taylor’s research group at Florida State University reported the atomic structure of the smooth muscle myosin molecules.
The similarity suggests, Padrón said, that the interacting head structure may be common to relaxed-state myosin for smooth and striated muscle and among varied species. “This model is applicable across the whole animal kingdom and all muscle types, and that’s exciting,” he remarked.
Padrón said he hopes to apply the research to muscle diseases that arise from malfunctioning of the muscles’ on/off switches. One such disease is hypertrophic cardiomyopathy, in which the wall of the left ventricle of the heart becomes enlarged, causing sudden death. It is caused by mutations in certain genes that encode several muscle proteins—some of which are related specifically to the myosin that Padrón studies.
Media Contact
More Information:
http://www.hhmi.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…





















