Venn diagram tactics to vet complex disease

NDUFS6 mutations are a novel cause of lethal neonatal mitochondrial complex I deficiency
A whole range of human muscular and neuromuscular diseases are caused by mutations in the mitochondrial respiratory chain/oxidative phosphorylation system. The problem is that there are about 120 genes involved in this system, some that are found in the mitochondria, and thus inherited through the mother, and some that are found in the nucleus and are inherited from both the mother and the father. Between the number of genes involved and the variable inheritance patterns, identifying all the mutations that could cause disease is a daunting task. Now David Thorburn and colleagues, from Murdoch Childrens Research Institute, have developed a methodology that takes a “Venn Diagram” approach to efficiently pinpoint mutations in this system. They utilize this method to identify a new cause for a lethal neonatal disease. The strategy makes use of cell-fusion experiments where cells from different patients are fused and the authors ask whether the fused cells are no longer defective — this means the cells can complement one another. If the fused cells complement each other, this means that the original cells had mutations in different genes.
The authors then carry out a second round of fusions, but in these cells, they have had the mitochondrial DNA removed. Here, if the fused cells are no longer defective, the mutation is in the patient’s nuclear DNA. If the fusions remain defective, the causal mutation is likely to be in the mitochondrial DNA. This technique is a very rapid and powerful means to categorize the mutation type of cells from different patients. The authors effectively demonstrated the strength of this technique using cells from 10 different patients, and showed that 7 contained nuclear mutations and 1 contained a mitochondrial mutation. The authors then focused on two patients whose cells did not complement one another, meaning the same gene was mutated. From there they used a battery of mapping, microarray and bioinformatics analyses and identified mutations for both patients in the NDUFS6 gene; a gene not previously shown to be involved in this type of disease. This work shows how combining a series of analyses, including genetic, biochemical, microarray, and bioinformatics techniques provide a rapid and effective means to pinpoint mutations in diseases that have complex genetic backgrounds.
Media Contact
More Information:
http://www.jci.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…
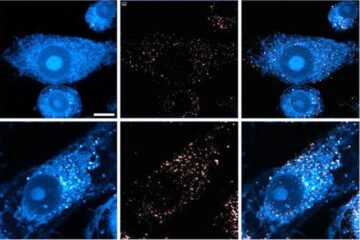
Innovative microscopy demystifies metabolism of Alzheimer’s
Researchers at UC San Diego have deployed state-of-the art imaging techniques to discover the metabolism driving Alzheimer’s disease; results suggest new treatment strategies. Alzheimer’s disease causes significant problems with memory,…
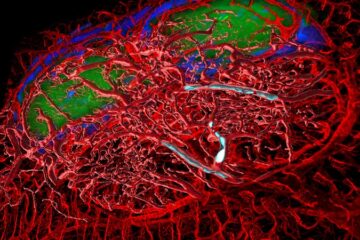
A cause of immunodeficiency identified
After stroke and heart attack: Every year, between 250,000 and 300,000 people in Germany suffer from a stroke or heart attack. These patients suffer immune disturbances and are very frequently…





















