Researchers identify the genome’s controlling elements

Scientists have churned out genome sequences for everything from fungi to dogs to chimps, and they won’t be letting up any time soon. However, because a genome sequence is little more than a static list of chemicals–like, say, a parts list for a 747 airplane–scientists are increasingly turning their attention to figuring out how living organisms put their genes to work. Using yeast as a testing ground, researchers at Whitehead Institute for Biomedical Research have for the first time revealed all the “controlling elements” of an entire genome–findings that may soon contribute to a new way of understanding human health and disease. “This is really the next stage in human genome research,” says Whitehead Member Richard Young, who headed the project together with Whitehead Fellow Ernest Fraenkel and MIT Computer Scientist David Gifford.
Key to understanding how the genome is controlled are gene regulators, also known as transcription factors. These small molecules intermittently land on a region of DNA, close to a particular gene, and then switch that gene on. They can also influence the amount of protein that the gene will produce. Many diseases, such as diabetes and cancer, are associated with mutated gene regulators, which is one reason why scientists are so interested in them.
The problem is that very few of these regulators have been identified in any organism. Locating their landing sites is essential to identifying their function, and therein lies the rub: Gene regulators are hard to find. They typically just land on a small stretch of DNA, do their job, and then take off again. And owing to the vastness of the genome, locating just one gene regulator with conventional lab tools can take many years. The Whitehead/MIT team, in the September 2 issue of the journal Nature, report developing a method for scanning an entire genome and quickly identifying the precise landing sites for these regulators.
This work builds upon research reported by Young in the journal Science in October 2002, in which he mapped the general locations of approximately half the gene regulators in yeast. “The results of the Science paper were pretty low-resolution,” says Young. “We were only able to identify in a general way regions where these gene regulators landed. In this paper, we’ve located all 203 regulators in yeast,” and using tools developed in Fraenkel’s lab have also been able to nail down the exact landing points. As a result, scientists now can begin to understand how genes and their regulators “talk” to each other. According to Fraenkel, knowing these communication patterns ultimately will have a profound influence on our understanding of everything from infectious disease to cloning.
To eavesdrop on these cellular conversations, graduate student Chris Harbison from Young’s lab and postdoctoral researcher Ben Gordon from Fraenkel’s lab combined the latest biological tools with new computational methods.
Harbison took yeast cells and subjected them all to a dozen nutritional, chemical, and temperature environments. “We tried to come up with different conditions that a yeast cell would encounter in its natural habitat,” says Harbison.
Gene regulators come out of hiding and do their job in response to environmental conditions, but they don’t all respond to the same kinds of predicaments. Running the cells through a wide spectrum of stimuli was a way of waking up all the regulators–in a sense, shaking the bushes and then nabbing them once they’re out.
Next, Harbison placed gene fragments associated with these regulators onto a series of microarrays–small dime-sized silicon or glass chips that contain thousands of pieces of DNA–which allowed him to come up with a list of approximate locations. Gordon and Fraenkel created computer algorithms that fused Harbison’s data with data from other yeast species in order to find the exact landing points. “The microarray information is like a noisy telephone call,” says Fraenkel. “We needed to find the words in all the static before we could understand what they meant.”
The next challenge is to scale the platform so it can tackle human cells, something that the researchers are gearing up to do. In fact, a paper from Young’s lab in the journal Science last February anticipated this possibility. The team reported locating in human cells all the genome-wide landing sites of a handful of gene regulators associated with type 2 diabetes, revealing some surprising mechanisms of the disease. “That paper is just the beginning of what we’ll soon be doing in human cells,” Young declares.
Even though the yeast genome’s 203 regulators are a far cry from the roughly 2,000 in human cells, Young explains, “now we have the technology and the concepts to get started on decoding the human genome.”
Media Contact
More Information:
http://www.wi.mit.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
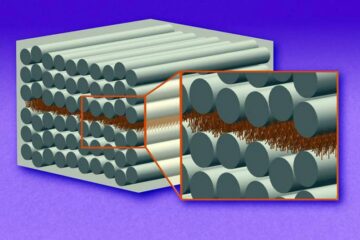

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
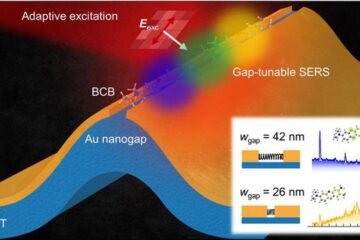
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















