Exploring small RNA function
Regulation of gene expression–deciding how much of what proteins are produced in the cell–is controlled by a myriad of different molecules. One type of naturally occurring regulatory molecule is small interfering RNA (siRNA), which selectively disrupts the production of a protein it is programmed to recognize, a process called RNA interference. These short stretches of nucleotides combine with other cellular proteins to form an RNA-induced silencing complex, called RISC, which locates and destroys a targeted messenger RNA–the molecule that carries a protein recipe from the nucleus to the site of production in the cytoplasm. Biologists have discovered hundreds of other naturally occurring short pieces of regulatory RNA, called microRNAs, in both plants and animals. Like siRNA, they also affect gene expression, through similar, possibly even identical RISC molecules. Animal microRNAs, however, target messenger RNA at a different stage in protein production. Though researchers have determined the sequences of these microRNAs, uncovering their function–that is, which protein they interrupt and, in turn, what the interrupted protein does–has progressed slowly and sporadically without any decisive tool to study them. Only four animal microRNAs have known biological functions, despite the intense level of work going on in this field.
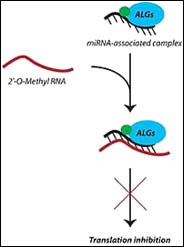
In order to determine the functions of siRNAs and microRNAs, Gyorgy Hutvagner and colleagues have developed a system whereby short stretches of 2’-O-methyl oligonucleotides whose sequence mirrors the targeted siRNA or microRNA, bind and inhibit their function, allowing researchers an unprecedented glimpse at the regulatory roles and mechanisms behind RNA interference. Hutvagner and colleagues constructed an oligonucleotide inhibitor based on the sequence of a microRNA called let-7, which blocks the production of the protein Lin-41 and is important for proper developmental timing in roundworm larvae. Larvae injected with the oligonucleotide had the exact features of a let-7 deficient worm, showing that the inhibitor did indeed block this microRNA’s function. Furthermore, the authors also used the oligonucleotides to provide evidence that two proteins, previously suggested to be involved with let-7, were directly associated with its interfering activity.
Using the technique described here, scientists could make rapid headway toward uncovering the biological functions of hundreds of microRNAs, their accessory RISC proteins, and even the proteins and genes they are programmed to interrupt. Furthermore, finding that RISC production is saturable could have significant implications for genetic studies that use RNA interference to uncover the function of sequenced, but unknown, genes; knowing the minimum required concentration of siRNA, researchers can avoid a buildup and any unwanted cell activity that goes along with it.
Citation: Hutvagner G, Simard MJ, Mello CC, Zamore PD (2004) Sequence-Specific Inhibition of Small RNA Function. PLoS Biol 2(4): e98 DOI: 10.1371/journal.pbio.0020098
CONTACT:
Phillip Zamore
University of Massachusetts Medical School
Worcester, MA 01605
United States of America
508-856-2191
phillip.zamore@umassmed.edu
This article is presented as a pre-issue publication. It will be part of our April 2004 issue
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…





















