Genome researcher analyze chromosome 7

New study discovers unusual structural features implicated in disease
A detailed analysis of the reference sequence of chromosome 7 has uncovered structural features that appear to promote genetic changes that can cause disease, researchers from the International Human Genome Sequencing Consortium said today.
In a study published in the July 10 issue of the journal Nature, a multi-institution team, led by the Washington University School of Medicine in St. Louis, reported it had sequenced 99.4 percent of the gene-containing region of chromosome 7 to an accuracy of greater than 99.99 percent. The team also described its analysis of this highly accurate reference sequence, an effort that took advantage of recently released data on the mouse genome to refine gene predictions and zero in on chromosomal regions that may be of special interest in understanding genetic diseases.
In addition to representing the largest chromosome to date to undergo detailed sequence analysis, chromosome 7 is significant because it has served as a pioneering chromosome for genomic and genetic studies. Researchers first developed genome mapping techniques on chromosome 7, and in the late 1980s, this chromosome was also the first to be searched by a then-novel technique called positional cloning in the successful hunt for the cystic fibrosis gene.
“Chromosome 7 has long been of interest to the medical community. Besides containing many genes that are crucial to development, this chromosome also holds the gene for cystic fibrosis and is frequently damaged in some types of leukemia and other cancers,” said Francis S. Collins, M.D., Ph.D., director of the National Human Genome Research Institute (NHGRI), which funded, and also participated in, the study. “This new analysis, coupled with our commitment to free and unrestricted access to sequence data, should further speed the discovery of genes on chromosome 7 related to human health and disease.”
Among the study’s most interesting results was the finding that, compared with previously analyzed human chromosomes, chromosome 7 contains an unusually high amount of duplicated sequence segments, covering roughly 8 percent of its DNA sequence. Researchers do not yet know the mechanism behind this high rate of duplication and also do not know why the duplication is much more extensive on the short arm of chromosome 7 than on its long arm.
However, in their study, researchers noted that this segmental duplication may encourage the type of genetic deletions that cause disease, as appears to be the case with the chromosomal region implicated in Williams-Beuren syndrome.
The syndrome, which is characterized by growth deficiency, heart disorders and mild mental retardation, is associated with very large deletions in a region of the long arm of chromosome 7 – a region that the new analysis also found to be a hotbed of duplicated segments. Based on previous, smaller-scale studies, genetic scientists know that such duplicated segments, or duplicons, serve to encourage large-scale deletions and other dramatic rearrangements of genetic material. It is also known that, in addition to their potential to cause disease by disrupting genes, such genetic rearrangements may on rare occasions be beneficial by facilitating the formation of new genes.
Richard K. Wilson, Ph.D., director of the Washington University School of Medicine’s Genome Sequencing Center and lead author of the study, said, “Our findings underscore the dynamic nature of the human genome and reveal how sequence structure may provide us with new insights into the genetic basis of human disease. But this analysis also drives home the fact that we still have a long way to go – that we are just taking our first steps down the pathway to understanding the complicated interplay of genomics and health. Each chromosome that we analyze will likely add a new twist or turn.”
In their analysis of the highly polished reference sequence, Dr.Wilson and his colleagues identified approximately 1,150 protein-coding genes on chromosome 7, about 20 percent less than the 1,455 predicted in a previous study by a different team.
The accuracy and completeness of the human chromosome 7 sequence assembled by the International Human Genome Sequencing Consortium was evaluated in part by Eric D. Green, M.D., Ph.D., and his colleagues at NHGRI’s Genome Technology Branch. When they compared the representation of markers called sequence-tagged sites (STSs) in the recently assembled sequence with STSs in previously constructed physical and genetic maps of chromosome 7, the NHGRI researchers found an excellent overall concordance.
Dr. Green, who is NHGRI’s scientific director and a co-author of the study, also emphasized the value of comparing the sequence of human chromosome 7 to its recently sequenced counterpart in the mouse. “Comparing the human sequence to the mouse sequence allowed our team to perform much more rigorous analyses of genes than would have otherwise been possible. The ability to place the human sequence alongside the mouse sequence helped us to swiftly distinguish real, protein-coding genes from pseudo-genes. The power of comparative genomics really sharpened our focus,” Dr. Green said.
In addition to NHGRI and Washington University, other institutions taking part in the chromosome 7 analysis were: University of Washington Genome Center, Seattle; University of California, Santa Cruz; Case Western Reserve University School of Medicine, Cleveland; and EMBL, Heidelberg, Germany.
There are 23 pairs of chromosomes in the human genome, which bear the 3 billion DNA letters that carry the genetic blueprint for human life. Chromosome 7 is one of the larger chromosomes, containing about 5 percent of the DNA in the human genome.
The sequencing work on chromosome 7 was carried out at the Genome Sequencing Center at the Washington University School of Medicine as part of the Human Genome Project, which receives substantial funding from NHGRI. The Human Genome Project officially began in October 1990 and was completed in April 2003. The entire project, including genetic mapping, technology development, the study of model organisms, and the ethical, legal and social implications (ELSI) program, was completed more than two years ahead of schedule at a cost that was $400 million less than expected.
The initial analysis of the draft human genome sequence was published in Nature in February 2001. With the completion of the Human Genome Project, researchers plan to publish a separate analysis on each completed chromosome over the next year or so. In addition to chromosome 7, researchers with the International Human Genome Sequencing Consortium have already published analyses of chromosomes 14, 20, 21, 22 and Y.
NHGRI is one of the 27 institutes and centers at the National Institutes of Health, an agency of the Department of Health and Human Services. Additional information about NHGRI can be found at its Web site, http://www.genome.gov.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
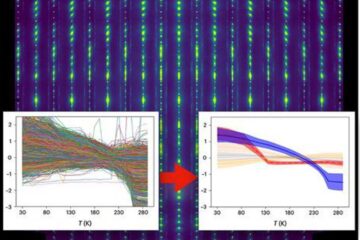
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
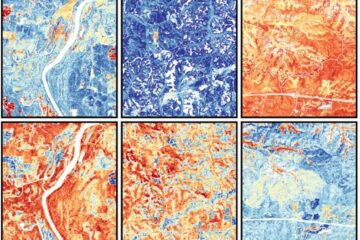

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















