FISH-BOL: NOAA Researchers Help Build a Global Reference Library of DNA Barcodes

Most of us are familiar with bar codes, those small black stripes with numbers below, known as the Universal Product Code or UPC label, that appear on commercial products. We scan them at the grocery store or to check a price, or have to cut them out and send them in for a rebate.
Now imagine scanning a DNA barcode on the piece of fish you just bought for dinner to instantly verify the species, where it came from, its nutritional value, and other valuable information. NOAA researchers are helping to make this scenario a reality.
“We need to accurately identify species for a number of reasons, from documenting the biodiversity of poorly sampled species and geographic areas to understanding populations and managing global fisheries in a sustainable way,” said Bruce Collette, a zoologist at NOAA’s National Systematics Laboratory (NSL) located in the Smithsonian Institution in Washington DC. “DNA barcoding is another tool in the toolbox of taxonomists and researchers who study, document, and organize knowledge about all life forms on earth.”
Collette and colleagues at NSL, part of NOAA’s Northeast Fisheries Science Center, are participating in FISH-BOL, the global Fish Barcode of Life Initiative, which plans to collect at least five representatives each of all 30,000 plus marine and freshwater species in the world. FISH-BOL is part of the global Consortium for the Barcode of Life (CBOL), started in 2003 to barcode everything from fishes, mushrooms and flowers, to microbes, insects and animals of every description.
With decades of experience, Collette is among a small number of taxonomists who conduct the painstaking work required to identify new species, or re-identify a specimen as new information is gained. After years looking into microscopes and examining animals stored in jars, he now spends much of his time working at a computer screen at his NOAA office deep within the Smithsonian’s National Museum of Natural History.
“Even though several million species of plants, animals and microbes have been identified over the past 300 years or so, we still find new species,” Collette says. “And despite advances in technology, we still have to sort organisms into piles and research every known bit of information before we can say we found a new species. Sometimes we have to go back to the basics and look at an organism in a jar for reference to make a determination.”
Scientists like Collette often go to sea and collect dozens of samples of an organism they think might be a new species. They return to the lab to sort out what they have collected before focusing in on the details to determine exactly what the organism is, sharing samples with colleagues to reach a consensus. The process takes time, and they are excited about the contributions FISH-BOL and other similar efforts around the world will make to documenting and understanding life forms on earth.
After decades of following similar taxonomic procedures often done by visual identification, DNA barcoding offers a new and much faster, more accurate way to identify species and share information. Since nearly all biological species have distinct gene sequences, they can be identified using a short gene sequence collected from a standardized position in the genome – a DNA barcode. Barcoding of animals relies on differences between species in a relatively short segment of mitochondrial DNA.
The first step in FISH-BOL and other similar efforts is to build a public library of barcode sequences from museum reference specimens. Specimens provide tissue samples that produce a reference barcode for that specific species. Researchers using standard tools and techniques in molecular biology extract DNA from the tissue, locate and isolate the barcode region, then replicate and sequence the genes. Then the specimens, known as voucher specimens, are preserved and catalogued into archival museum collections.
Not only can specimens be identified as part of a known species using only a tiny piece of tissue from the organism, but new variations in what was thought to be a single species can be determined. Tissue from unidentified specimens can also be matched to the DNA sequence of a known species in the reference library, a process that currently takes just a few hours and a few dollars but soon may take only minutes and cost pennies.
Barcode records of each specimen contain the DNA sequence, information about the voucher or referenced specimen, and the species name. The records are stored in three global databases and are available without charge. As an electronic database, FISH-BOL contains DNA barcodes, images and geographic information of examined specimens, as well as linkages to the voucher specimens, information on species distributions, nomenclature, taxonomic information, natural history information and literature citations.
“Barcoding works for all stages in the life cycle, so it will help us identify larval fish, which is timely since many leaders in larval fish taxonomy are retiring,” Collette said. “It can also differentiate between closely related species that are hard to tell apart, especially large specimens that are difficult to bring back from the field. And it can positively identify fishery products like fish fillets so you know if the grouper you ordered in a restaurant is really grouper.”
For fishery managers and researchers, barcoding can legally verify identifications of fishes caught as by-catch and species under regulation, important to protecting endangered species and sustaining fish populations.
Collette and researchers at NSL have already started adding to the voucher specimens at the National Museum of Natural History by barcoding fishes in the Gulf of Maine and western North Atlantic. So far, tissues have been collected from 508 specimens representing 162 species, including 101 species from the Gulf of Maine out of the 252 known. He will present some of the group’s results at the July 2008 meeting of the American Society of Ichthyologists and Herpetologists in Montreal, Canada.
Media Contact
More Information:
http://www.noaa.govAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
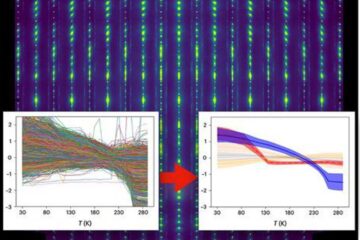
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
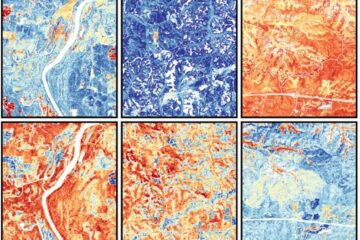

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















