Omics analysis nurtures the creation of functional plants

Masami Yokota Hirai
Team Leader
Metabolic Systems Research Team
Metabolomics Research Division
RIKEN Plant Science Center
Plants produce a wide variety of metabolites from inorganic compounds, some with useful functions including health-promoting effects. The ability to harness these metabolites by creating ‘functional plants’ that produce these compounds in large quantities could therefore be of considerable benefit to society. It is also an intriguing research topic for plant scientists. Progress in this field relies on our understanding of how the hundreds of thousands of unique metabolite compounds are produced in plants. Masami Yokota Hirai at the RIKEN Plant Science Center (PSC) is investigating the mechanism of metabolism in plants by linking metabolites to genes through omics analysis—a combination of metabolome and transcriptome analyses. The knowledge obtained from this approach is leading steadily to the successful development of functional plants.
Compound synthesis by metabolism in plants
“The word ‘metabolism’ may remind many people of a reaction in which the proteins or lipids we consume as food are degraded and converted into energy. Metabolism in plants, however, is different from this reaction,” says Masami Yokota Hirai, team leader of the Metabolic Systems Research Team at the RIKEN Plant Science Center (PSC). “Metabolism in plants is a reaction in which inorganic compounds such as nitrogen, phosphorus, and sulfur are absorbed through the roots and light energy is used to produce various organic compounds including amino acids and sugars such as starch. These organic compounds are called metabolites.”
The production of compounds such as amino acids, sugars and vitamins, which make up the body of a plant, from inorganic compounds is called ‘primary metabolism’, whereas the production of more complex compounds from primary metabolites is called ‘secondary metabolism’. “A plant is rooted in place and cannot move. To cope with environmental hazards, such as insects, dry weather or salt damage, plants produce secondary metabolites as they are exposed to these stresses. A plant maintains a constant amount of primary metabolites, but produces secondary metabolites on an as-needed basis.”
Hirai’s focus on metabolism in plants is motivated by the potential uses of the vast array of metabolites that plants produce. “Plants produce metabolites that specific to the plant species. There are more than 200,000 known unique metabolites, some of which contain nutrient, health-promoting and medical ingredients.” Examples of secondary metabolites include isoflavone, anthocyanin, menthol, catechin and capsaicin (Fig. 1). These compounds are drawing attention for their health-promoting functions. “Efficient production of useful secondary metabolites will greatly help improve our health. For this reason, we are working hard to elucidate the metabolic mechanism in plants.”
Metabolome studies
“A metabolic map illustrates how metabolites are produced from inorganic substances through specific reactions, and can be as complicated as a subway route map. A partial metabolic map does not provide enough information to understand the whole picture of metabolism. We need to understand the metabolic system as a whole. To help with this, we have started to take advantage of omics analysis.”
Omics is a science that comprehensively embraces the four disciplines of genomics, transcriptomics, proteomics and metabolomics (Fig. 2), and has developed rapidly in the past decade triggered by improvements in genome decoding techniques and processing speed. These improvements have led to some remarkable milestones in genomic research, including sequencing of the complete genomes of the flowering plant Arabidopsis thaliana, rice and soy-bean.
Similarly, the DNA microarray technique used in transcriptome analysis, the analysis of RNA transcribed from DNA, has also improved dramatically over the past decade. In this technique, hundreds of thousands of single-stranded DNA fragments are fixed in holes or ‘spots’ on a glass substrate, and fluorescently labeled RNA are dropped onto the substrate surface. RNA complementary to a DNA fragment will become bound to the DNA, which causes the combined compound to emit fluorescent light. From the fluorescence intensity of each spot, researchers can determine which genes are being expressed and to what extent. The expression of Arabidopsis genes has been analyzed using this method and the data has been made publicly available via the AtGenExpress database.
Progress in metabolome analysis, however, lags considerably behind that of genomic and transcriptome analyses. “Metabolome analysis is the least developed part of omics,” says Hirai. “In metabolome analysis, a mass spectrometer is used to determine the mass of molecules and electrical charges contained in a specimen. From this data we can determine the types and quantities of metabolites in a specimen. The work, however, is extremely difficult. Genomics focuses only on DNA and transcriptomics on RNA, and the same method can be used for all types of organism. In metabolomics, on the other hand, we deal with metabolites with a wide range of characteristics, such as volatility and water solubility, which makes it impossible to conduct investigations under fixed conditions. The metabolites are also produced in highly variable amounts, tiny to large quantities, and with a wide range of concentrations. So we need to share a single specimen among a number of measuring instruments so that enough data can be collected.”
The DNA microarray technique is almost entirely automated, which allows almost anybody to use it, whereas mass spectroscopy requires a highly skilled operator. This has obstructed rapid developments in metabolome analysis. Another factor lies with the collected data itself. “The data obtained by mass spectrometry are plotted on a graph with mass along the horizontal axis and intensity along the vertical axis. A peak in the graph corresponds to a single metabolite. Of the thousand or so peaks that are produced, only about 10% have been assigned to specific metabolites. Most of the metabolites remain unknown. We are at a loss regarding where to begin,” says Hirai.
Research into metabolomics started in 2000. In Japan, Kazuki Saito, group director of the Metabolomic Function Research Group at the PSC, took the initiative in research on metabolomics. In those days, Saito was a professor at the Graduate School of Pharmaceutical Sciences at Chiba University, and Hirai attended his laboratory. “I was confident that the research was not only interesting but also very important. However, I had a hard time for about three years because I could not find a way to understand the data itself,” says Hirai, looking back on those days.
The world’s first omics analysis
In 2004, Hirai published a paper on metabolome analysis in the Proceedings of the National Academy of Sciences, USA. The article would become the most-cited paper in the field of plant biotechnology in 2005. “This paper is a collection of results obtained from an integrated analysis of the transcriptome and metabolome of A. thaliana. The paper does not provide new information on gene functions or metabolite synthesis, so I am not completely satisfied with the paper because it is a simple description of my research results. But in those days, few papers could be found on metabolome analysis. I think that the paper was highly evaluated under such circumstances because I tried to derive something new by combining transcriptome analysis with metabolome analysis for Arabidopsis. It was a pioneering attempt at omics analysis.”
Investigation of all 27,000 Arabidopsis genes shows that there are multiple genes that express with the same timing. These genes are likely to be involved in the same function. If the population of a metabolite increases while a certain gene cluster is expressing and decreases when the gene cluster is not expressing, the metabolite could be associated with the gene cluster. In this way, omics analysis allows genes to be linked with metabolites, making it possible to understand metabolite function.
Creating vegetables with cancer-preventing effects
Arabidopsis synthesizes about 30 kinds of glucosinolates from methionine and tryptophan, which can be degraded by myrosinase (an enzyme) into isothiocyanate (a ‘spicy’ flavor component). Sulforaphane is a kind of methionine-derived isothiocyanate and induces the production of phase-2 enzymes that detoxify carcinogens. The gene PMG1 has a stimulating effect only on the synthesis of glucosinolates from methionine.
One successful application of omics analysis is the 2007 discovery of a new gene that makes cruciferous vegetables produce cancer-preventing components. Cruciferous vegetables such as broccoli, radish, horse radish and mustard have a ‘spicy’ flavor that has been attributed to pungent components that originate as metabolites called glucosinolates, among which sulforaphane is known to enhance the functions of enzymes that detoxify carcinogens. Using Arabidopsis, a member of the cruciferous family, Hirai successfully showed that the gene PMG1 controls the synthesis of glucosinolates.
“Through an integrated analysis of the transcriptome and metabolome of Arabidopsis, we found a gene cluster that changed with the same pattern. The gene cluster was found to contain the genes that are known to be involved in the synthesis of glucosinolates. In this way, we looked into the genes, and finally reached PMG1.”
It was also confirmed that the functional enhancement of PMG1 in Arabidopsis promotes glucosinolate synthesis. “As the amount of glucosinolate increases, the amount of sulforaphane increases. This could allow us to grow vegetables with enhanced cancer-preventing effects.”
A study focusing on gene clusters associated with the synthesis of glucosinolates is now under way. Hirai is interested in the gene BASS5. The base sequence of BASS5 is similar to that of the genes for bile acid transporter, an animal protein. Bile acid transporter is present in the cell membrane and is responsible for the intercellular movement of bile acids. When the base sequences are similar, their functions are also often similar. But since there are no bile acids in plants, the role of the proteins created by BASS5 is particularly interesting. “The proteins were thought at first to be present in the cell membrane where they mediate the intercellular movement of glucosinolates. However, we have come to understand that the proteins are likely to be present not in the cell membrane but on the surface of the cell’s chloroplast. This demonstrates that secondary metabolites are synthesized not only in the cellular cytoplasm, but also in the chloroplasts. We think BASS5 might be associated with glucosinolate intermediates moving in and out of the chloroplasts.” By omics analysis it was confirmed that glucosinolates are not created when BASS5 function is inhibited. Omics analysis has been demonstrated in this way time and time again to be a very effective tool for elucidating metabolic pathways.
Hirai is also researching the synthesis of amino acids, particularly methionine, the primary metabolite from which glucosinolates are produced. “An increase in a metabolite requires an increase in the supply of its source material, namely amino acids. We should know how to synthesize amino acids if we are to attempt to create plants that produce large amounts of useful secondary metabolites.”
Metabolome analysis in full swing
“Metabolome analysis will proceed rapidly in the years to come,” says Hirai with confidence. This is thanks to an analytical technique called widely targeted metabolome analysis developed by Yuji Sawada, a special research scientist in the Metabolic Systems Research Team. Associating the thousands of peaks produced by mass spectrometry with metabolites has been one of the obstacles to metabolome research. “In conventional targeted metabolomics, observations are made with the aim of identifying a single known kind of metabolite,” says Hirai. “Widely targeted metabolomics is based on the idea that if the number of target metabolites can be increased, eventually all metabolites could be targeted, which could lead to full metabolome analysis. In widely targeted metabolomics, all of the peaks in the data correspond to known metabolites. This allows us to proceed to the next stage of the research program immediately. Usually, analytical methods are developed by analytical chemists and information scientists, but Dr Sawada and myself are biologists. Widely targeted metabolomics is a very convenient analytical method for biologists.” Widely targeted metabolomics currently handles about 700 target metabolites, a number that will be increased in the near future.
Hirai also hopes to develop a new analytical method for omics analysis. “It is essential to develop additional tools if we are to remain pioneers in this field because integrated analysis of transcriptome and metabolome data is now available to everyone. We are now working on developing, by trial and error, a new analytical method that can suggest unforeseen results.”
The eureka moment, the best part of being a researcher
“The analytical transcriptome and metabolome data are a description of the state of a plant. Using the data to understand the biological implications depends on our ability as scientists, so I feel the data is testing us,” says Hirai. “Sometimes, an idea flashes through my mind when I look at the same data for hours or even days, which could lead to a discovery. I find it interesting to look back over my laboratory notebook to find exclamation marks written next to certain notes. I believe I had little eureka moments when I wrote those exclamation marks. Those moments are the best part of being a researcher.”
About the Researcher
Masami Yokota Hirai
Masami Yokota Hirai was born in Chiba, Japan, in 1965. She graduated from the Department of Agricultural Chemistry in the Faculty of Agriculture at The University of Tokyo in 1989, and received her PhD from the Graduate School of Agriculture at the same institution in 1994. She moved to RIKEN in 2005 after working as a research associate in the Graduate School of Pharmaceutical Sciences at Chiba University. She has been in her current position since 2008 as team leader of the Metabolic Systems Research Team at the RIKEN Plant Science Center.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

New SPECT/CT technique shows impressive biomarker identification
…offers increased access for prostate cancer patients. A novel SPECT/CT acquisition method can accurately detect radiopharmaceutical biodistribution in a convenient manner for prostate cancer patients, opening the door for more…

How 3D printers can give robots a soft touch
Soft skin coverings and touch sensors have emerged as a promising feature for robots that are both safer and more intuitive for human interaction, but they are expensive and difficult…
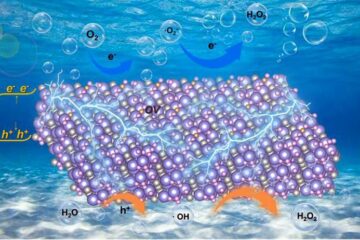
Oxygen vacancies mediated ultrathin Bi4O5Br2 nanosheets
… as efficient piezocatalyst for synthesis of H2O2 from pure water. As an important chemical raw material, hydrogen peroxide (H2O2) is widely applied in various aspects of industry and life….





















