Mapping translation sites in the human genome

While the general mechanism of translation has been understood for some time, protein synthesis can initiate by more than one mechanism. One of the least well understood mechanisms is known as cap-independent translation.
Now, John Chaput and his colleagues at Arizona State University's Biodesign Institute have produced the first genome-wide investigation of cap-independent translation, identifying thousands of mRNA sequences that act as Translation Enhancing Elements (TEEs), which are RNA sequences upstream of the coding region that help recruit the ribosome to the translation start site.
The new study outlines a technique for mining whole genomes for sequences that initiate cap-independent translation within the vastness of the genome.
The research has important implications for the fundamental understanding of translation in living systems, as well as intriguing potential in the biomedical arena. (Many viral pathogens are known to use cap-independent translation to hijack and redirect cellular mechanisms to translate viral proteins.)
The lead author of the study is Brian P. Wellensiek, a senior scientist in Biodesign's Center for Evolutionary Medicine and Informatics. The group's results appear in the current issue of the journal Nature Methods.
During most protein synthesis in eukaryotic cells, cap-dependent translation dominates. The process begins after DNA is first transcribed into mRNA, with the aid of an enzyme polymerase. mRNA now forms the coded template from which the translated proteins will be generated. The mRNA code consists of sequences made from 4 nucleic acids, A, C, G & U, with each 3-letter grouping (known as a codon), corresponding to one amino acid in the protein being synthesized.
A key component in the translation process is the ribosome, which migrates along the single stranded mRNA, reading the codons as it goes. Before it can do this however, it must locate a special structure at the 5' end of the mRNA strand known as the cap. In normal cap-dependent translation, the ribosome is recruited to the 5' end of mRNA via a specialized cap-binding complex.
Cap-independent translation allows the ribosome to begin reading the mRNA message without having to first locate the 5' cap structure. Cap-independent translation occurs in eukaryotic cells during normal processes including mitosis and apoptosis (or programmed cell death). It is also a feature in many forms of viral translation, where the viral transcript is able to recruit the ribosome and co-opt its function to preferentially translate viral RNA.
In the current study, Chaput designed an in vitro selection strategy to identify human genome sequences that initiate cap-independent translation. The technique is able to select candidates from a pool of trillions of genomic fragments. Once a set of sequences was identified as translation enhancing elements, they were shown to function effectively in both cell-free and cellular translation systems.
As Chaput explains, most research on cap-independent translation has been conducted using RNA fragments derived from viruses. “These RNA molecules will fold into shapes that appear to mimic some of the initiation factors that that you would find in eukaryotic translation,” he says. More recently, similar RNA molecules have been identified in cellular systems, though the sequences tend to be much shorter and function in a different manner.
Chaput's method of studying such sequences on a genome-wide scale involves first generating a DNA library of the entire human genome. Using enzymes, the genome is cut into random fragments of around 200 base pairs each. These sequences are then transcribed into mRNA.
Applying a technique known as mRNA display, the fragments are tagged in specific way, such that amino acid sequences resulting from successful translation events remain bound to the mRNA fragments that generated them. “Essentially, what we're doing is taking a library of human mRNA and tagging those sequences that act as translation enhancing elements,” Chaput says. Those sequences bearing an attached peptide affinity tag can then be separated out from the remaining untranslated sequences, reverse transcribed, amplified using PCR technology and subjected to subsequent rounds of selection.
The sequences were later mapped onto the human genome. As expected, the complete library of sequences used at the start of the experiments mapped fairly evenly across the genome. But the sequences selected via mRNA display as translation enhancing elements tended to cluster in non-coding regions of the genome. The authors speculate that such sequences may have been evolutionarily selected against, as they have the potential to disrupt normal cap-dependent translation.
Roughly 20 percent of the translation enhancing elements functioned as internal ribosomal initiation sites, again turning up primarily in non-coding genomic regions. The origin of these sequences remains mysterious. It is conceivable that they were surreptitiously brought on board as a result of human interaction with different types of viruses.
Once Chaput's group had acquired a library of 250 distinct translation enhancing elements through selection using mRNA display, the sequences were screened for translation enhancing activity, which was quantified using a light based assay employing a luciferase reporter molecule.
By measuring levels of luciferase, the enhancement of each sequence could be assessed relative to background noise, with the better translation enhancing elements displaying 50-100 fold enhancement (and some as much as 1000-fold enhancement). The next step was to determine which of these sequences could function as internal ribosomal initiation sites.
To do this, the same 250 sequences were inserted into a vector bearing a hairpin structure. As Chaput explains: “If the ribosome latched onto the 5' end, it would hit that hairpin and would fall off. However if the ribosome skipped the hairpin and recognized the sequence on the other side of the hairpin independently and translated it, that's an indication that the sequence is functioning as an internal ribosomal initiation site.” Both assays (for translation enhancement and internal ribosomal initiation) were validated under cell-free conditions and in human cells, using a vaccinia virus vector.
A study of this scope is possible thanks to innovative techniques for in vitro selection (such as mRNA display), as well as a revolutionary technology permitting massively parallel RNA sequencing (known as deep sequencing), which provides unprecedented speed and read accuracy.
Much remains to be learned about atypical translation processes. The mechanism of action for translation enhancing elements is still obscure, particularly in the case of internal ribosomal initiation sites. Similarly, the particular gene products that may result from cap-independent translation have yet to be identified and characterized.
The research group consisted of:
Brian P. Wellensiek1, Andrew C. Larsen1, Bret Stephens1, Kim Kukurba1,2, Karl Waern3, Natalia Briones1, Li Liu1, Michael Snyder3, Bertram L. Jacobs4,5, Sudhir Kumar1,5 and John C. Chaput1,2*
1Center for Evolutionary Medicine and Informatics in the Biodesign Institute, Arizona State University, Tempe, AZ 85287, 2Department of Chemistry and Biochemistry at ASU, 3Center for Genomics and Personalized Medicine, Department of Genetics, Stanford University, Stanford, CA 94305, 4Center for Infectious Diseases and Vaccinology in the Biodesign Institute at ASU, 5School of Life Sciences at ASU.
Written by:
Richard Harth
Science Writer: Biodesign Institute
Media Contact
More Information:
http://www.asu.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
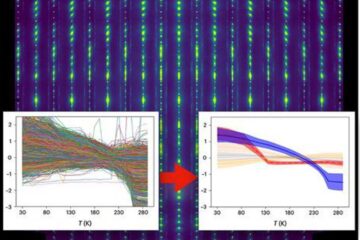
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
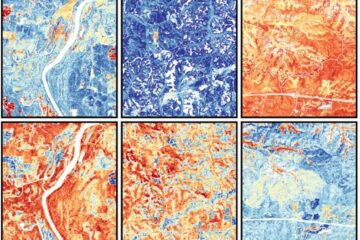

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















