Investigators Discover New Gene That Affects Clearance of Hepatitis C Virus
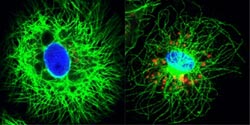
The novel protein, IFNL4—stained in red—as expressed in primary human liver cells treated to mimic the effect of infection with Hepatitis C (left). The protein shows up only in carriers of the unfavorable allele (right). <br>
They also identified an inherited variant within this gene, Interferon Lambda 4 (IFNL4), that predicts how people respond to treatment for hepatitis C infection. The results of this study, by investigators at the National Cancer Institute (NCI), part of the NIH, and their collaborators at NIH and other institutions, were published online in Nature Genetics on Jan. 6, 2013.
Chronic infection with hepatitis C virus is a cause of liver cirrhosis and liver cancer. Up to 80 percent of people who are acutely infected with hepatitis C fail to clear the virus and develop chronic hepatitis C infection, and of these, approximately 5 percent develop liver cancer. Individuals of African ancestry do not respond as well to current treatments of hepatitis C infection compared to patients of European or Asian ancestry.
Previously, results from genome-wide association studies (GWAS) identified common inherited genetic markers that were associated with response to hepatitis C virus treatment and spontaneous clearance of the infection. Those markers are located on chromosome 19 near a known interferon gene, IFNL3 (IL28B). However, molecular investigations into IFNL3 did not explain the GWAS association with spontaneous virus clearance or treatment response. To find the new gene, the investigators used a technology involving RNA sequencing on human liver cells treated to mimic hepatitis C virus infection.
“By using RNA sequencing we looked outside the box to search for something beyond what was already known in this region. We hit the jackpot with the discovery of a new gene. It is possible that other important genes may be discovered using this approach,” said co-lead investigator Ludmila Prokunina-Olsson, Ph.D., of the Laboratory of Translational Genomics in NCI’s Division of Cancer Epidemiology and Genetics (DCEG).
The researchers found that the IFNL4 region harbors a variant that is found in two alternative forms. One form, called deltaG, results in a deletion in one of the four bases that comprise DNA. The change creates an alteration known as a frameshift, which produces the IFNL4 protein, while the form without the deletion does not produce IFNL4. By analyzing data from hepatitis C-infected African-Americans and European-Americans participating in clinical studies, the authors found that the presence of the IFNL4 protein is associated with poorer clearance and response to treatment than the form that does not produce IFNL4. The deletion variant is more common in people of African ancestry, which helps partially explain why African-Americans have a lower response to current hepatitis C treatments than patients of Asian and European ancestry.
“Our work fulfills several promises of the genomic era,” said NCI’s Thomas R. O’Brien, M.D., Infections and Immunoepidemiology Branch, DCEG. “One, a better understanding of biology; two, personalized medicine; and three, new potential treatments. We deliver immediately on the first two. We’ve identified a new gene that may help us better understand response to viral infection and the new genetic marker may transition to clinical practice because it predicts treatment outcome for African-American patients better than the current genetic test. For the third, the INFL4 protein may be used as a novel therapeutic target for hepatitis C virus infection, and possibly other diseases.”
The new gene belongs to what is now a family of four interferon-lambda protein-encoding genes, three of which were discovered more than ten years ago (IFNL1, IFNL2 and IFNL3) The mechanism by which the IFNL 4 protein impairs hepatitis C virus clearance remains unknown. Further studies will explore molecular function of this novel protein in normal and disease conditions.
This study was conducted collaboratively with the National Institute of Diabetes and Digestive and Kidney Diseases at NIH, as well as the U.S. Food and Drug Administration, and a number of universities and research institutions. Funding was provided by NCI grant Z01 CP005782.
Media Contact
More Information:
http://www.nih.govAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















