First 3D Model of a Synapse
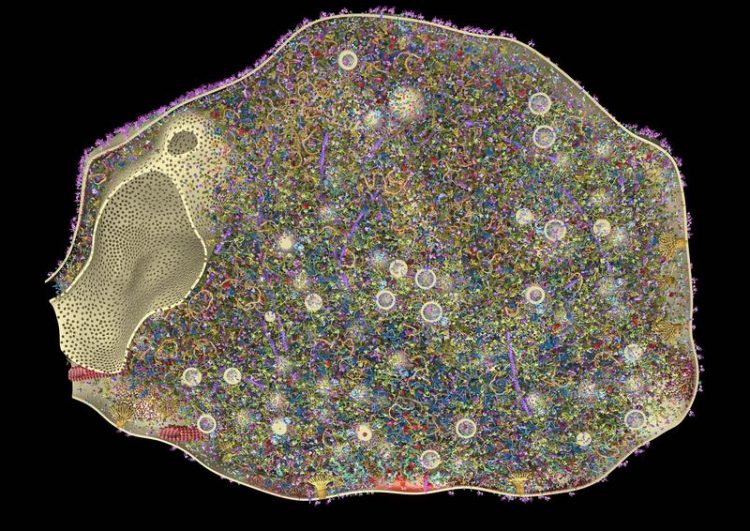
The first 3D model of a synapse. The reconstruction shows a synapse in cross section. The small spheres are synaptic vesicles. The model shows 60 different proteins, which sum to more than 300,000 copies of a single protein in the synapse. Burkhard Rammner
Synapses are the contacts between nerve cells that allow the flow of information that makes our brains work. However, the molecular architecture of these highly complex structures has been unknown until now. A research team from Göttingen, led by Prof. Silvio O. Rizzoli from the DFG Research Center and Cluster of Excellence Nanoscale Microscopy and Molecular Physiology of the Brain (CNMPB) of the University Medical Center Göttingen, managed to determine the copy numbers and positions of all important building blocks of a synapse for the first time.
This allowed them to reconstruct the first scientifically accurate 3D model of a synapse. This effort has been made possible only by a collaboration of specialists in electron microscopy, super-resolution light microscopy (STED), mass spectrometry, and quantitative biochemistry from the UMG, the Max Planck Institute for Biophysical Chemistry, Göttingen, and the Leibniz Institute for Molecular Pharmacology, Berlin.
The project was funded by the European Research Council (ERC) and the Deutsche Forschungsgemeinschaft (DFG). The results have been published in the renowned scientific journal SCIENCE on May 30th 2014. Highlighting the impact of this work, the presented model has been selected as the cover of the respective issue of the SCIENCE journal.
Original publication:
Wilhelm BG, Mandad S, Truckenbrodt S, Kröhnert K, Schäfer C, Rammner B, Koo SJ, Claßen GA, Krauss M, Haucke V, Urlaub H, Rizzoli SO (2014) Composition of isolated synaptic boutons reveals the amounts of vesicle trafficking proteins. SCIENCE, May 30th, 2014. 344(6187): 1023-1028. DOI: 10.1126/science.1252884.
A New and Undiscovered Land in Neuroscience
“This 3D model of a synapse opens a new world for neuroscientists,” says Prof. Rizzoli, senior author of the publication. Particularly the abundance and distribution of the building blocks have long been terra incognita, an undiscovered land. The model presented by Prof. Rizzoli and his team now shows several hundreds of thousands of individual proteins in correct copy numbers and at their exact localisation within the nerve cell.
“The new model shows, for the first time, that widely different numbers of proteins are needed for the different processes occurring in the synapse,” says Dr. Benjamin G. Wilhelm, first author of the publication. The new findings reveal: proteins involved in the release of messenger substances (neurotransmitters) from so called synaptic vesicles are present in up to 26,000 copies per synapse. Proteins involved in the opposite process, the recycling of synaptic vesicles, on the other hand, are present in only 1,000-4,000 copies per synapse.
These details help to solve a long-lasting controversy in neuroscience: how many synaptic vesicles within the synapse can be used simultaneously? Apparently, more than enough proteins are present to ensure vesicle release, but the proteins for vesicle recycling are sufficient for only 7-11% of all vesicles in the synapse. This means that the majority of vesicles in the synapse cannot be used simultaneously.
The most important insight the new model reveals, is however that the copy numbers of proteins involved in the same process scale to an astonishingly high degree. The building blocks of the cell are tightly coordinated to fit together in number, comparable to a highly efficient machinery. This is a very surprising finding and it remains entirely unclear how the cell manages to coordinate the copy numbers of proteins involved in the same process so closely.
The new model will serve as a reference source for neuroscientists of all specialisations in the future, and will support future research, since the copy number of proteins can be an important indicator for their relevance. But the research team led by Prof. Rizzoli does not plan to stop there: “Our ultimate goal is to reconstruct an entire nerve cell”. Combined with functional studies on the interaction of individual proteins this would allow to simulate cellular function in the future – the creation of a “virtual cell”.
An impressive video animation has been created from the obtained data to visualise the structure and protein distribution of a synapse (http://www.sciencemag.org/content/344/6187/1023/suppl/DC1).
For his approach to study the molecular anatomy of nerve cells, Prof. Rizzoli has already been awarded an ERC Consolidator Grant 2013. “The findings of Prof. Rizzoli are spectacular”, says Prof. Dr. Heyo Kroemer, Speaker of the UMG board and dean of the Faculty of Medicine. “This highly precise synaptic model will provide completely new possibilities for medical research. This is another example demonstrating that the University Medical Center Göttingen provides attractive conditions for international top-level research.” Prof. Mathias Bähr, one of the CNMPB speakers, says “We are very excited that Prof. Rizzoli succeeded in accomplishing this project so fast. This study significantly advances our general understanding of the protein distribution within healthy nerve cells. In the future, these findings will contribute to identify anomalies in neuronal anatomy in neurodegenerative diseases, such as Parkinson’s disease.”
FURTHER INFORMATION:
Prof. Dr. Silvio O. Rizzoli
Dept. Neuro- and Sensory Physiology, University Medical Center Göttingen European Neuroscience Institute (ENI) Göttingen
Grisebachstraße 5, 37077 Göttingen
Phone: +49 (0) 551 39-33630
srizzol@gwdg.de
CNMPB – Center for Nanoscale Microscopy and Molecular Physiology of the Brain Cluster of Excellence 171 – DFG Research Center 103
Dr. Heike Conrad
Scientific Coordination / Press & Public Relations
Humboldtallee 23, 37073 Göttingen
Telefon 0551 / 39-7065, heike.benecke@med.uni-goettingen.de
http://RMATION:
http://www.rizzoli-lab.de – research group of Prof. Rizzoli
http://www.cnmpb.de – DFG Research Center and Cluster of
http://Excellence Nanoscale Microscopy and Molecular Physiology of the Brain (CNMPB) –
http://www.sciencemag.org/content/344/6187/1023/suppl/DC1 – video animation
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…





















