Efficient working in confined spaces: New insights into the architecture of cellular protein factories
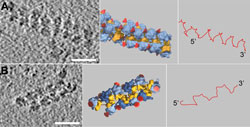

Figure: The three dimensional strucuture of the polysome:Cryoelectron tomographic picture of two polysomes (left), schematic diagram of their structure (middle) and the messenger molecule (mRNA) pathway within the polysome (right). The small ribosomal subunits (yellow) are oriented towards the inside of the polysome, the large subunits (blue) and the nascent protein chains (indicated by red cones) face the cytosol. If the ribosomes are continuously arranged in a \"top-to-top\" orientation (Fig. A, middle), the result is a pseudo-helical structure of the polysome. If the ribosomes are arranged alternating in \"top-to-top\" and in \"top-to-bottom\" orientation (Fig. B, middle), the result is a staggered structure. In both cases the mRNA traverses the shortest possible path from one ribosome to its next neighbor (Fig A and B, right). Florian Brandt, Max-Planck-Institut für Biochemie<br>
In order to generate as many proteins as possible at the same time, several ribosomes cluster together to form an “industrial complex” – the polysome – and read simultaneously the same messenger molecule. Scientists at the Max-Planck-Institute of Biochemistry have now, for the first time, been able to reveal the three-dimensional structure of these complexes (Cell 23.1.2009).
In a polysome, the ribosomes are densely packed and exhibit preferred orientations: The small ribosomal subunits are orientated towards the inside of the polysome and the ribosomes are arranged either in a staggered or in a pseudo-helical structure (see figure). This arrangement ensures that the distance between nascent protein chains is maximized, thereby reducing the probability of intermolecular interactions that would give rise to aggregation and limit productive folding. Until now, the belief has been that specialised proteins, the so-called chaperones, would prevent protein misfolding.
Against the background of the new findings, their function appears in a new light: “It appears possible that the main function of chaperones that interact with nascent polypeptide chains is not to suppress chain aggregation within polysomes, but rather to reduce intra-chain misfolding as well as aggregation between different polysomes in the crowded cellular environment”, explains Ulrich Hartl, head of the “Cellular Biochemistry” department, who lead the project in cooperation with Wolfgang Baumeister, head of the “Molecular Structural Biology” department.
Moreover, the spatial structure of the polysome enables the ribosomes to process the messenger molecule in the protected area within the polysome and to pass it on without detours. Thus, the architecture of the cellular protein factories facilitates an optimized work flow and increases the efficiency of protein folding.
Original Publication:
The Native 3D Organization of Bacterial Polysomes; Florian Brandt, Adrian H. Elcock, Stephanie A. Etchells, Julio O. Ortiz, F. Ulrich Hartl and Wolfgang Baumeister;
Cell, DOI 10.1016/j.cell.2008.11.016
Contact:
Florian Brandt
Max-Planck Institut für Biochemie
Am Klopferspitz 18
D-82152 Martinsried
Germany
mail: fbrandt@biochem.mpg.de
Dr. Monika Gödde
Public Relations
Max-Planck-Institut für Biochemie
Am Klopferspitz 18
82152 Martinsried
phone: 089 – 8578 3882
mail: goedde@biochem.mpg.de
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
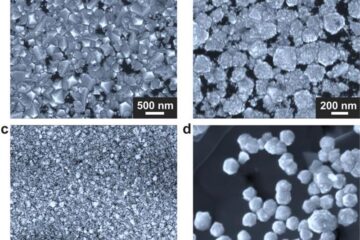

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…





















