Dark Matter Made Visible Before the Final Cut

The new study reveals snippets of information contained in dark matter that can alter the way a gene is assembled.
“These small sequences of genetic information tell the gene how to splice, either by enhancing the splicing process or inhibiting it. The research opens the door for studying the dark matter of genes. And it helps us further understand how mutations or polymorphisms affect the functions of any gene,” said study senior author, Zefeng Wang, PhD, assistant professor of pharmacology in the UNC School of Medicine and a member of UNC Lineberger Comprehensive Cancer Center.
The study is described in a report published in the January 2013 issue of the journal Nature Structural & Molecular Biology.
The findings may be viewed in terms of the film industry’s editorial process where snippets of celluloid are spliced, while others end up unused on the proverbial cutting room floor.
Taken from a DNA point of view, not every piece of it in each human gene encodes for a functional protein; only about 10 percent does, in coding regions called “exons.” The other 90 percent of the stuff that fills the intervening regions are longer stretches of dark matter known as “introns.”
But something mysterious happens to introns during the final processing of messenger RNA (mRNA), the genetic blueprint that’s sent from the cell’s nucleus to its protein factory. Only particular exons may be included within the final mRNA produced from that gene, whereas the introns are cut out and destroyed.
It’s therefore easier to understand why more scientific attention has been given to exons. “When people are looking at the genetics of a disease, most of the time they’re looking for the change in the coding sequence,” Wang said. “But 90 percent of the sequence is hidden in the gene’s introns. So when you study gene variants or polymorphisms that cause human disease, you can only explain the part that’s in the exon. Yet the majority remains unexplainable because they’re in the introns.”
Following completion of the genome sequencing projects, subsequent DNA and RNA sequencing revealed the existence of more than one splice variant, or isoform, for 90 percent of human genes. During messenger RNA processing, most human genes are directed to be cut and pasted into different functional isoforms.
In a process called alternative splicing, a single gene could code for multiple proteins with different biological functions. In this way, alternative splicing allows the human genome to direct the synthesis of many more proteins than would be expected from its 20,000 protein-coding genes.
“And those different versions sometimes function differently or in opposite ways,” Wang said. “This is a tightly regulated process, and a great number of human diseases are caused by the ‘misregulation’ of splicing in which the gene was not cut and pasted correctly.”
Wang’s research colleagues identified “intronic splicing regulatory elements.” These essentially recruit protein factors that can either enhance or inhibit the splicing process.
Their discovery was accomplished by inserting an intron into a green fluorescent protein (GFP) “reporter” gene. These introns of the reporter gene carried random DNA sequences. When the reporter is screened and shows green it means that portion of the intron is spliced.
“The default is dark,” so any splicing enhancer or silencer can turn it green,” Wang explains. “In this unbiased way we can recover hundreds of sequences of inhibited or enhanced splicing.”
The study collaborators put together a library of cells that contain the GFP reporter with the random sequence inserted. Thus, when researchers looking at the intron try to determine what a particular snippet of genetic information does and its effect on gene function, they can refer to the splicing regulatory library of enhancers or silencers.
“So it turns out that the sequencing element in both exons and introns can regulate the splicing process, Wang says. “We call it the splicing code, which is the information that tells the cell to splice one way or the other. And now we can look at these variant DNA sequences in the intron to see if they really affect splicing, or change the coding pattern of the exon and, as a result, protein function.”
Collaborators in this study with Zefeng Wang are Yang Wang in the department of pharmacology and member of the UNC Lineberger Comprehensive Cancer Center; Meng Ma, Anhui University, Hefei, China; and Xinshu Xiao, University of California, Los Angeles.
Support for the research comes from the American Heart Association and the National Cancer Institute, a component of the National Institutes of Health.
Media Contact
More Information:
http://www.unc.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
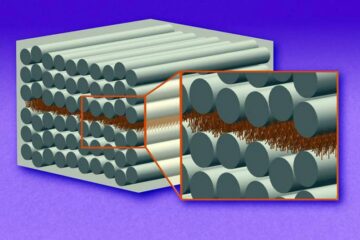

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
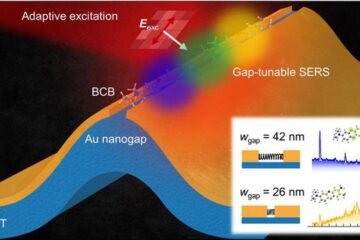
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















