Bacterial proteins: A structural switch leads to multifunctionality in gene expression
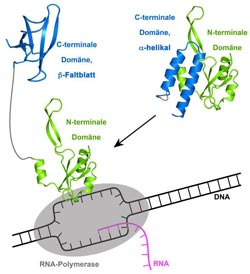
In the closed form of RfaH (right), the C-terminal domain (CTD, blue) and the N-terminal domain (NTD, green) are close to each other. The alpha-helical CTD masks the area of the NTD which binds to the RNA polymerase. Binding to a specific piece of DNA results in domain separation (left) which in turn leads to the complete refolding of the CTD. In this state, the NTD can bind RNA polymerase and the CTD can bind ribosomal protein S10. RfaH is thus a regulatory component of the transcription of DNA into RNA.<br><br>Image: Dr. Stefan Knauer, University of Bayreuth; free for publication only when reference is included.<br>
The bacterial protein RfaH is able to adopt two completely different three-dimensional structures. Effected by external factors, the carboxyterminal domain switches from an all alpha helical to an all beta barrel conformation. This drastic conformational change enables the regulation of gene expression and protein translation by RfaH.
Nuclear magnetic resonance spectroscopy reveals the extraordinary structural switch of a protein
Proteins, basic molecular building blocks of life, consist of a chain of amino acids which usually adopts a unique three-dimensional structure dictated by the sequence of amino acids. Most proteins can fulfill specific functions only in a folded state. The traditional scientific view states that in a defined environment a certain protein can adopt only one distinct three-dimensional structure to accomplish its purpose.
Recent results from the Research Center for Bio-Macromolecules at the Universität Bayreuth evidenced that this view has to be modified: The protein RfaH from E. coli bacteria was studied in an international cooperation led by Prof. Paul Rösch. Using nuclear magnetic resonance spectroscopy, the researchers could show that the bacterial protein RfaH is able to adopt two completely different three-dimensional structures. Results of bacterial genetics studies demonstrate that the two structures accomplish entirely different functions. RfaH consists of two characteristic structural units, the aminoterminal domain (N-terminal domain, NTD) and the carboxyterminal domain (C-terminal domain, CTD), that are connected by a flexible linker. These domains are closely interacting and are thus in close proximity to each other. The CTD consists solely of two alpha helices (screw-like structures) in a hairpin arrangement. Binding of the NTD to a distinct piece of DNA leads to spatial separation of the domains, which in turn results in a complete structural switch of the CTD as it changes its structure from the alpha helical hairpin into a fold that completely differs from the starting structure (beta sheet).
“Never before has such a fundamental structural change been observed for proteins”, Prof. Paul Rösch notes. “This result is spectacular as we were able to simultaneously elucidate the structural transition and its functional consequences for central cellular processes in bacteria.” RfaH's ability to change its structure enables regulation of the translation of bacterial genetic information into proteins (gene expression).
Regulatory functions of the domains in gene expression
Gene expression starts with the transcription of the genetic information contained in DNA into RNA. The molecular machine for this process is RNA polymerase. Transcription is followed by the production of new proteins based on RNA (translation) at a different cellular component, the ribosome.
Its ability to change its structure allows RfaH to physically couple the main actors of these processes. After domain dissociation, the NTD of the protein binds to RNA polymerase, while the refolded CTD interacts with the ribosome. This binding is mediated by the protein S10 that is part of the ribosome. The spectacular structural switch of RfaH enables the protein to couple transcription and translation by bridging the two principal components, RNA polymerase and ribosome. The option of a regulated domain separation and the resulting complete refolding of the CTD explains the central role of the protein RfaH in the modulation of bacterial gene expression on the level of molecular structures.
The partner proteins RfaH and NusG
Why does RfaH exist in a non-functional state at all? Studies of the protein NusG provided hints to an answer. NusG like RfaH consists of an NTD and a CTD, but the two domains are always separated, and the CTD exists in beta sheet structure only. The NTD of NusG also binds to RNA polymerase and the CTD to the ribosome via S10. However, NusG is a protein that is generally involved in bacterial gene expression while, in contrast, RfaH is employed only in very specialized transcription events. To ensure that RfaH does not interfere with NusG, the alpha helical CTD of RfaH masks precisely the area of the NTD which could interact with RNA polymerase. Also, the CTD in its alpha helical state is not able to bind to the ribosome. Thus, RfaH is inhibited in both functions. The protein is activated by domain separation and refolding of the CTD to beta sheet structure – only then the two domains can bind their partners.
International cooperation
These results published in “Cell” are the outcome of a long-standing transatlantic cooperation. The Research Center for Bio-Macromolecules (BIOmac) at the Universität Bayreuth led by Prof. Paul Rösch has cooperated closely with biochemists, bacteriologists, and microbiologists of Ohio State University and of the University of Wisconsin. The Deutsche Forschungsgemeinschaft (DFG) in Germany and the National Institutes of Health (NIH) in the USA supported the research.
Outlook
“Together we discovered an example of how a protein can change its fold fundamentally to be able to fulfill different functions”, Prof. Paul Rösch explains. “The principle of making proteins multifunctional by switching their three-dimensional structure is so strikingly simple that we are prepared to find similar mechanisms in other molecular processes.”
Publication:
Burmann et al., An α Helix to β Barrel Domain Switch Transforms the Transcription Factor RfaH into a Translation Factor,
Cell (2012), http://dx.doi.org/10.1016/j.cell.2012.05.042
Svetlov and Nudler, Unfolding the Bridge between Transcription and Translation,
Cell (2012), http://dx.doi.org/10.1016/j.cell.2012.06.025
Burmann et al., A NusE:NusG Complex Links Transcription and Translation.
Science. 2010 328:501-4.
Video:
An explanatory video can be found at http://www.cell.com
Contact:
Prof. Dr. Paul Rösch
Forschungszentrum für Bio-Makromoleküle
Universität Bayreuth
D-95440 Bayreuth
Tel. +49 (0)921 55-3540
E-Mail: roesch@unibt.de
Media Contact
More Information:
http://www.uni-bayreuth.deAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…





















