A new way to extract bone-making cells from fat tissue
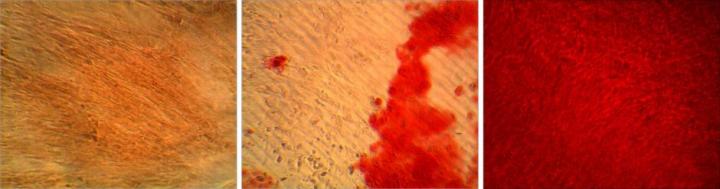
Finding cells that have bone-making potential is more efficiently done by looking at the genes they express -- ALPL, in this case -- than at proteins on their surface. The bone matrix being produced by cells is stained red in samples of cells that are sorted for not expressing ALPL (left), sorted for expressing ALPL (right), and left as an unsorted control (center). Credit: Darling lab/Brown University
In a new study in the journal Stem Cell Research & Therapy, scientists at Brown University demonstrate a new method for extracting a wide variety of potential bone-producing cells from human fat. They developed a fluorescent tag that could find and identify cells expressing a gene called ALPL. Expression of the gene is an indicator of bone-making potential.
If the tag finds the RNA produced when the gene is expressed, it latches on and glows. A machine that detects the fluorescing light then separates out the ALPL-expressing cells.
In the paper, the scientists report that their method produced more than twice the yield of potential bone-makers (9 percent) compared to their best application of another method: sorting cells based on surface proteins presumed to indicate that a cell is a stem cell (4 percent).
Brown University has applied for a patent on the method of gene expression tagging for producing a tissue.
Meanwhile, the ALPL-expressing cells produced on average more than twice as much bone matrix (and as much as nine times more in some trials) during three weeks of subsequent cultivation than a similar-sized population of unsorted adipose tissue cells and almost four times more bone matrix than cells that don't express ALPL. ALPL-expressing cells were also better at making cartilage or fat.
A couple of other research groups have also sorted stem cells based on gene expression, but they have not done so specifically with the goal of enriching cell populations for a specific tissue, the researchers said.
Lead author and Brown graduate student Hetal Marble said targeting gene expression rather than surface proteins for the purpose of gathering cells to make a new tissue is a “paradigm shift” in the following regard: Gene expression provides a way to target any cell based on whether it can produce another tissue, while targeting surface proteins limits researchers to harvesting cells that fit a presumed definition of being a stem cell. The new approach, she said, is more pragmatic for the purpose.
“Approaches like this allow us to isolate all the cells that are capable of doing what we want, whether they fit the archetype of what a stem cell is or not,” Marble said. “The paradigm shift is thinking about isolating populations that are able to achieve an end point rather than isolating populations that fit a strictly defined archetype.”
In their experiments, though, the team tolerated a four-day delay that they'd like to dispense with in the future. It takes that long for the maximum number of cells to express ALPL when cells are chemically primed to do so.
In future research, said senior author Eric Darling, the Manning Assistant Professor of Molecular Pharmacology, Physiology and Biotechnology and a member of the Center for Biomedical Engineering assistant professor of medical science, the team would like to target a gene expressed much earlier in the differentiation process to see if they can avoid a priming period.
If they can apply the method based on a gene that's expressible within a matter of hours, that could allow future surgeons working on bone healing to take out some of a patient's fat cells, sort out the best bone-producers (primed or not) and then implant those cells in the bone break within the same surgical session.
“If you can take the patient into the OR, isolate a bunch of their cells, sort them and put them back in that's ideally where we'd like to go with this,” Darling said. “Theoretically we could do this with other genes that might upregulate very quickly or are innately expressed.
###
In addition to Marble and Darling, other authors are Bryan Sutermaster, Manisha Kanthilal, and Vera Fonseca.
The National Science Foundation (CBET1253189), The National Institutes of Health (R01 AR063642, P20 GM104937) and the U.S. Department of Education (P200A120064) provided support for the study.
Media Contact
More Information:
http://www.brown.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















