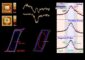
Using living cells as nanotechnology factories

The results were published in the early online edition of the Proceedings of the National Academy of Sciences.
Yan specializes in a fast-growing field within nanotechnology — commonly known as structural DNA nanotechnology — that uses the basic chemical units of DNA, abbreviated as C, T, A, or G, to self-fold into a number of different building blocks that can further self-assemble into patterned structures.
“This is a good example of artificial nanostructures that can be replicated using the machineries in live cells” said Yan. “Cells are really good at making copies of double stranded DNA and we have used the cell like a copier machine to produce many, many copies of complex DNA nanostructures.”
DNA nanotechnologists have made some very exciting achievements during the past five to 10 years. But DNA nanotechnology has been limited by the need to chemically synthesize all of the material from scratch. To date, it has strictly been a test tube science, where researchers have developed many toolboxes for making different DNA nanostructures to attach and organize other molecules including nanoparticles and other biomolecules.
“If you need to make a single gram of a DNA nanostructure, you need to order one gram of the starting DNA materials. Scientists have previously used chemical methods to copy branched DNA structures, and there has also been significant work in using long-stranded DNA sequences replicated from cells or phage viruses to scaffold short helper DNA sequences to form 2-D or 3-D objects,” said Yan, who is also a professor in the Department of Chemistry and Biochemistry at ASU.
“We have always dreamed of scaling up DNA nanotechnology. One way to scale that it up is to use the cellular system because simple DNA can be replicated inside the cell. We wanted to know if the cell's copy machine could tolerate single stranded DNA nanostructures that contain complicated secondary structures.”
To test the nanoscale manufacturing capabilities of cells, Yan and his fellow researchers, Chenxiang Lin, Sherri Rinker and Yan Liu at ASU and their collaborators Ned Seeman and Xing Wang at New York University went back to reproducing the very first branched nanostructure made up of DNA- a cross-shaped, four-arm DNA junction and another DNA junction structure containing a different crossover topology.
To copy these branched DNA nanostructures inside a living cell, the ASU and NYU research team first shipped the cargo inside a bacteria cell. They cut and pasted the DNA necessary to make these structures into a phagemid, a virus-like particle that infects a bacteria cell. Once inside the cell, the phagemid used the cell just like a photocopier machine to reproduce millions of copies of the DNA. By theoretically starting with just a single phagemid infection, and a single milliliter of cultured cells, Yan found that the cells could churn out trillions of the DNA junction nanostructures.
The DNA nanostructures produced in the cells were also found to fold correctly, just like the previously built test tube structures. According to Yan, the results also proved the key existence of the DNA nanostructures during the cell's routine DNA replication and division cycles. “When a DNA nanostructure gets replicated, it does exist and can survive the complicated cellular machinery. And it looks like the cell can tolerate this kind of structure and still do its job. It's amazing,” said Yan.
Yan acknowledges that this is just the first step, but foresees there are many interesting DNA variations to consider next. “The fact that the natural cellular machinery can tolerate artificial DNA objects is quite intriguing, and we don't know what the limit is yet.”
Yan's group may be able to change and evolve DNA nanostructures and devices using the cellular system and the technology may also open up some possibilities for synthetic biology applications.
“I'm very excited about the future of DNA nanotechnology, but there is a lot of work to be done. An interesting research topic to pursue is the interface of DNA nanostructures with live cells; it is full of opportunities,” said Yan.
Media Contact
More Information:
http://www.asu.eduAll latest news from the category: Materials Sciences
Materials management deals with the research, development, manufacturing and processing of raw and industrial materials. Key aspects here are biological and medical issues, which play an increasingly important role in this field.
innovations-report offers in-depth articles related to the development and application of materials and the structure and properties of new materials.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…